Getting a handle on neuronal behaviour in culture
Posted: 20 May 2005 | | No comments yet
In drug discovery for CNS diseases, the use of complex neural cell culture systems offers many advantages. Innovations in high content screening enable us to identify compounds which affect key cell biological properties in such cultures. We can bridge the divide between kinetic and endpoint screening by use of another novel technology, RNAi. High content […]
In drug discovery for CNS diseases, the use of complex neural cell culture systems offers many advantages. Innovations in high content screening enable us to identify compounds which affect key cell biological properties in such cultures. We can bridge the divide between kinetic and endpoint screening by use of another novel technology, RNAi.
High content screening (HCS) enables the detection of multiple types of individual morphological and biochemical responses in cell culture systems. Traditional drug discovery approaches to CNS drug discovery have involved measuring effects on kinases, GPCRs and ion channels, by use of plate readers, radioligand binding and first-generation imaging systems (FLIPR, VIPR, etc). Whilst these have proved entirely appropriate for many drug discovery targets, they are limited by being population measurements with an inherent lack of cellular or subcellular resolution. HCS has now begun to provide a greater level of resolution within pharmaceutical screening. However, first-generation high content screens have sometimes claimed to be novel, without actually changing drug discovery paradigms: that is, to be going beyond an additional way of detecting ion channel, or GPCR responses, rather to be generating information previously not attainable within the screening environment.
Innovative implementation of newer types of cell biology imaging within screening should enable drug discovery companies to target not so much an individually identified protein, already tractable for drug discovery, but rather to focus on the desired cellular event and potentially identify novel pathways leading to it. The ability to identify compounds that affect key cell biological properties e.g. kinetic signals, changes in cell morphology, or translocation of a protein of interest from one subcellular compartment to another, will be increasingly important in target and lead identification. Automated quantitation of complex cellular events should enable entirely new types of cellular and biochemical screening cascades to be developed.
There is now a variety of high content instrumentation available for endpoint and kinetic, cell biology analysis. Instruments in widespread use for endpoint assays include GE IN Cell 1000 and Cellomics ArrayScan and at the higher end of the market the confocal Evotec Opera and IN Cell 3000 systems (Zemanová et al., 2003). A combination of confocality, tip-changeable liquid handling and flexibility in imaging wavelengths, make BD Pathway Bioimager (formerly Atto PathwayHT) an additional choice, particularly for kinetic high content screening (Zemanová et al., 2003).
Neural cell cultures
There are a variety of aspects of diseases that can be modelled in a cell culture system of appropriate complexity and design. One example would be the loss of synapses that contributes in vivo to progressive cognitive impairment. Substantial evidence suggests that synapses are defective prior to structural deterioration in neurological diseases – decreased presynaptic protein levels are seen in post-mortem brains (Rapoport, 2003) and cytosolic Ca2+ homeostasis may also be defective. Preparations available to study such events include primary neuronal cultures from rat or mouse, acute brain slices, organotypic cultures and neural stem cells from embryonic or foetal derived sources.
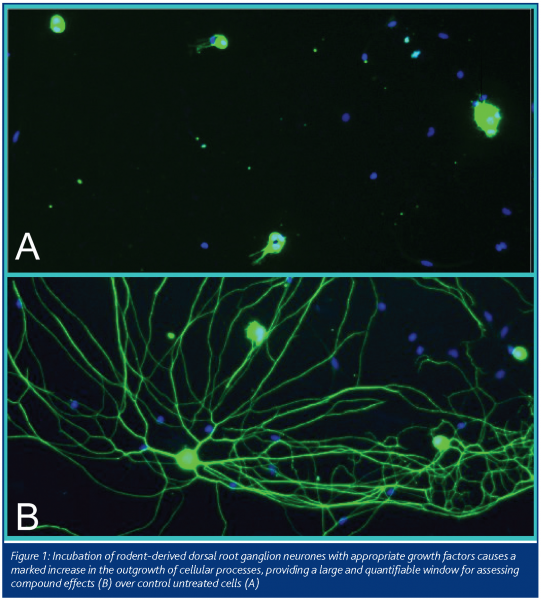
Cellomics ArrayScan instruments were the first to make available a commercial algorithm for an assay that is now commonplace across high content screening systems – quantification of neurite outgrowth. Niell et al (2004) have shown that synapses predominantly form on new neurites, such that neurite outgrowth may be regarded as a pre-requisite for formation of new synapses. One could view neurite outgrowth, therefore, as a surrogate assay for synapse formation, as well as being of biological interest in its own right. The first paper on use of this HCS approach came from my lab several years ago, (Simpson et al., 2001), validating the approach for neuroblastoma cells. Using cell subtype-specific antibodies, we and others have since shown that this can be readily adopted within mixed populations of cells, such as primary neuronal cultures or neural progenitor cell cultures (Figure 1).
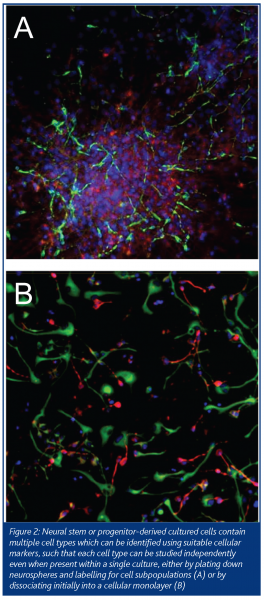
Neural stem or progenitor cells are an attractive drug discovery resource for multiple reasons, including the potential to screen on non-transformed human cells of the desired type to be targeted in the CNS. They usually grow and are passaged as neurospheres, with multiple cell types interacting. Culture conditions can be designed so that cells attach to the culture dish or microtitre plate as a monolayer or a neurosphere, producing a complex environment of cell types (progenitors, immature neurons, several types of glia) (Figure 2) – potentially the same interactions that are present within the developing intact CNS. At the current state-of-the-art these may present cellular variability between wells, which can be problematic for conventional drug discovery approaches. Using endpoint HCS, however, with appropriate subtype-selective antibodies, it is possible to overcome this issue by drilling down onto cell subpopulations and even to compare the effects of a compound on multiple subpopulations in the same well simultaneously. This can provide powerful information – for example, compounds causing changes within all cell types in the culture can be discarded in preference for compounds with selective effects on a subpopulation of interest.
Multiple HCS assay types
As it is never advisable to look at any single cellular event in isolation from measurement of other potential changes, it is attractive to be able to study the effects of an interesting compound on multiple cell biological events in parallel. Analysis of cell biology events can now be performed using a combination of technologies: ion channel recording; biochemical and molecular profiling; novel higher throughput electrophysiology techniques; kinetic high content assays (e.g. single cell video rate calcium or membrane potential imaging) and endpoint high content assays for synaptic physiology. All these assays can be appropriate for small molecule screening, or can be implemented in combination with RNAi technology to identify novel, molecular targets for drug discovery.
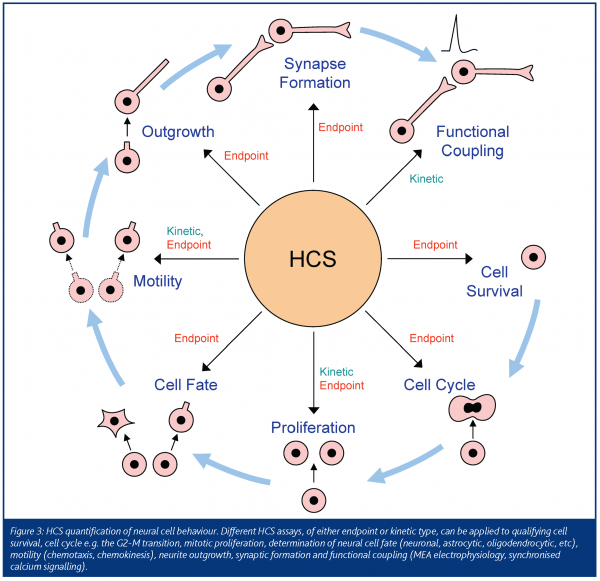
There are many events within the life cycle of a neural cell that can be quantified using HCS (Figure 3). Ourselves and others, have validated in CNS-derived cell cultures many approaches, including automated quantitation of effects on cell viability (live/dead, apoptosis), cell proliferation, cell motility (chemokinesis, chemotaxis) and outgrowth (Ramos Villulas et al., 2003; Ramm et al., 2003; Richards et al., 2004). Further detail is obtained by measuring different aspects of effects on each stage of the cell life cycle e.g. comparison of compound effects on G2-M transition cf G1 phase in cell cycle assays. Similarly, differential effects of stimuli on chemotaxis and chemokinesis, can be studied using variations of conventional motility assays e.g. Boyden chambers (e.g. Richards et al., 2004). Thereby, growth factors or compounds that are efficacious in one cellular event can be tracked across other related assays and the specificity of their effects assessed.
Multiplexed assays can be performed in the same well e.g., high-content kinetic assays can be followed by fixation and high-content endpoint characterisation, or multiple high-content algorithmic analyses performed on the same image sets. In addition, as only the cells – not the supernatant – are required for high-content imaging, it is possible to combine high-content assays with some conventional high throughput assays. Combining assays on the same cells and wells reduces inter-experimental variability and increases confidence in the separation of an effect of compound on one biochemical or cellular parameter over undesired effects on another parameter.
Kinetic HCS
A great deal of information can be gained from a single well in endpoint assays, however in a kinetic recording mode, this information is multiplied. For example, changes in calcium levels within cytosol or organelles may be tracked for minutes, or even hours, while studying >100 cells in each well of a microtitre plate. As the industry drives forward into new areas of assay design, there will be a need for data analysis, as well as image storage, to catch up. Simple quantitation of amplitude and rate of rise of responses will not be sufficient to capture the complexity of kinetic responses and communication from one cell to another, to ensure we fully understand the dynamic interactions within a complex cellular environment. Current commercial analysis tools from instrument vendors lack flexibility currently; however there are a number of customised approaches based on, for example, pattern analysis, from other biological arenas that are undoubtedly transferable into this rapidly developing area.
Small-scale integration with automation makes kinetic assays convenient for screening. In one example, at Merck Sharp and Dohme we have integrated a BD Pathway Bioimager with a CRS automated arm, incubator and PerkinElmer PlateTrak liquid handling workstation. This enables dye loading, cell incubation, cell imaging and posthoc cell fixing and staining, to integrate an entire kinetic drug discovery assay. One example of this assay type is to add a stimulator of calcium signalling to a population of mixed neural cells, track the calcium elevation in each cell, move the plates from the imager and fix and label for subtype-selective markers (e.g. nestin for neural precursors), then return plates to the imager and identify which cell types express the calcium elevation differentially. GFP-coupled intracellular proteins can make combined functional and morphological studies even easier. We have also needed to develop proprietary approaches for analysing the complex kinetic data that is generated using such assays.
Intracellular ionic changes can be intrinsic, as well as in response to extrinsic factors: the frequency of spontaneous oscillatory changes in intracellular calcium levels in network cultures correlates with synaptic density and other physiological changes (e.g. Cuddon et al., 2003). If these oscillations are not synchronised within a culture, or occur in only a minor subpopulation of cells, the activity would be missed by FLIPR/VIPR type population recording. Conventional electrophysiological recording of a large number of neurons simultaneously would also not be possible. However, either kinetic high content screening fluorescence systems, or multi-electrode array (MEA)-type extracellular electrophysiological recording devices, now permit information about intrinsic signalling to be quantified on a cell by cell basis. MEA biochips can be used to stimulate and record extracellular electrical activity of excitable biological tissue from as many as 60 recording sites simultaneously. Using multi-electrode, or imaging-based, approaches the temporal change in cellular responses at the onset of synchronicity can now be studied in great detail.
It is possible to bridge the divide between kinetic high content screening and endpoint high content screening using another technology, RNAi. By knocking down the level of expression of a single protein, one can examine the effects on a cell signalling response and the effect on change in cell phenotype, enabling the demonstration of a mechanistic link between biochemical and cell biological changes.
For changes in cellular responses, which take minutes or hours rather than seconds, there are other new options: reporter systems can be used to enable translocation events to be tracked using kinetic imaging, more conveniently than several predetermined time points checked using fixed cells on an endpoint imaging system. A variety of GFP-coupled protein constructs and cell lines for cellular biology investigation of e.g. translocation changes are available now for high content screening, from commercial vendors (e.g. Almholt et al., 2004). Long-term studies of ionic changes are also now possible with the new generation of detection technology, for example calcium sensitive dyes such as Fluo-LoJo or KoJo that do not so readily leach from cells or photo bleach.
Where next?
In summary, within the last couple of years powerful new HCS tools (confocal technology, new dyes and reporters and advances in stem and progenitor cell culture) have begun to enable studies not previously practical. This enables drug discovery scientists to evaluate and implement entirely new paradigms to identify targets and leads. What we may now regard as ‘conventional’ endpoint HCS is well complemented by kinetic HCS (cell imaging or electrophysiological approaches). Drug discovery scientists can now screen libraries of compounds or RNAi using relevant, more complex cell populations, e.g. primary neurones or progenitor cell-derived cultures, and generate a full set of functional data from cellular subpopulations. Long-term signalling mechanism studies using new dyes, reporters and confocal HCS technologies, in combination with array-based profiling tools, are available to help tease out the mechanism and molecular target of action, of active compounds. Finally, there is the attractive ability to combine function and phenotype studies at the same time in a mixed neural cell population, e.g. assays for ionic activity and cellular phenotype changes, using automation, to release the maximum potential of kinetic HCS assay design.
Acknowledgements
I am grateful to members of the Automated Imaging group who contributed to the experiments described in this review, and participated in discussions. I also thank the Internet and visual communications group for their assistance with Figure 3.



References
- Zemanová, L., Schenk, A., Valler, M.J., Nienhaus, G.U., & Heilker R. (2003) Confocal optics microscopy for biochemical and cellular high-throughput screening Drug Discovery Today 8, 1085-1093
- Rapoport, S.I. (2003) Coupled reductions in brain oxidative phosphorylation and synaptic function can be quantified and staged in the course of Alzheimer disease. Neurotox Res 5, 385-98
- Niell, C.M., Meyer, M.P. & Smith, S.J. (2004) In vivo imaging of synapse formation on a growing dendritic arbor. Nature Neurosci. 7, 254 – 260
- Simpson P.B., Bacha J.I., Palfreyman E.L., Woollacott A.J., McKernan, R.M., Kerby J. (2001) Retinoic acid-evoked differentiation of neuroblastoma cells predominates over growth factor stimulation: an automated image capture and quantitation approach to neuritogenesis. Anal. Biochem. 298, 163-169
- Ramos Villullas I., Smith A.L., Heavens R.P., Simpson P.B. (2003) Characterisation of a sphingosine 1-phosphate activated Ca2+ signalling pathway in human neuroblastoma cells. J. Neurosci. Res. 73, 215-226
- Ramm, P., Alexandrov, Y., Cholewinski, A., Cybuch, Y., Nadon, R., & Soltys, B.J. (2003) Automated screening of neurite outgrowth. J. Biomolecular Screening 8, 7-18
- Richards G.R., Millard R.M., Leveridge M., Kerby J.E., Simpson P.B. (2004) Quantitative assays of chemotaxis and chemokinesis for human neural cells. Assay Drug Dev. Technol. 2, 465-472
- Cuddon P., Roderick H.L., Simpson P.B., Berridge M.J., Bootman M.D. (2003) Spontaneous calcium oscillations control the development of metabotropic signalling in hippocampal neurons J Physiol. 547P
- Almholt D.L.C., Loechel F., Nielsen S.J.,Krog-Jensen C., Terry R., Bjørn S.P., Pedersen H.C., Præstegaard M., Møller S., Heide M., Pagliaro L., Mason A.J., Butcher S. and Dahl S.J. (2004) Nuclear export inhibitors and kinase inhibitors identified using a MAPK-activated protein kinase 2 Redistribution® screen. Assay Drug Dev. Technol. 2, 7-20




