RNAi therapeutics for neurodegenerative disease: challenges and prospects
Posted: 19 March 2008 | Matthew JA Wood, Department of Physiology, Anatomy and Genetics, University of Oxford | No comments yet
The early 21st century has seen a revolution in RNA biology, bringing with it the prospect of a new class of medicines based on RNA. What are the prospects for developing these RNA-based medicines for the growing medical problem of neurodegenerative disease and what are the challenges to making these new medicines work successfully within the complex environment of the nervous system? Recent progress on RNA silencing of neurodegenerative disease targets and RNAi delivery to the nervous system is encouraging and suggests that clinical evaluation of these therapeutic agents is realistic within the next few years.
The early 21st century has seen a revolution in RNA biology, bringing with it the prospect of a new class of medicines based on RNA. What are the prospects for developing these RNA-based medicines for the growing medical problem of neurodegenerative disease and what are the challenges to making these new medicines work successfully within the complex environment of the nervous system? Recent progress on RNA silencing of neurodegenerative disease targets and RNAi delivery to the nervous system is encouraging and suggests that clinical evaluation of these therapeutic agents is realistic within the next few years.
The early 21st century has seen a revolution in RNA biology, bringing with it the prospect of a new class of medicines based on RNA. What are the prospects for developing these RNA-based medicines for the growing medical problem of neurodegenerative disease and what are the challenges to making these new medicines work successfully within the complex environment of the nervous system? Recent progress on RNA silencing of neurodegenerative disease targets and RNAi delivery to the nervous system is encouraging and suggests that clinical evaluation of these therapeutic agents is realistic within the next few years.
A spectacular revolution in RNA biology over the last decade has created new tools and opportunities for advancing basic biomedical science as well as the tantalising prospect of RNA-based medicines to treat human disease. The discovery of RNA interference (RNAi) in 1998 by Fire and Mello1 led rapidly to elucidating the biochemical mechanism underlying the phenomenon of RNA-based gene silencing.
The key effector molecules were found to be small double-stranded RNAs (dsRNAs)2, the importance of these molecules at the centre of cellular gene regulatory networks is now appreciated. These small dsRNAs include exogenously derived small interfering RNAs (siRNAs) or endogenously expressed RNAs, for example, microRNAs (miRNAs). The mechanism of action of these dsRNAs, elucidated through biochemical and genetic experiments in a variety of organisms, involves processing (where necessary) by the Dicer complex to yield siRNAs, which associate with Argonaute 2 (AGO2) and the RNA-induced silencing complex (RISC). siRNA strand selection is governed by the thermodynamic properties of the duplex such that a guide strand is selected to direct target mRNA cleavage, which is carried out by the AGO2 catalytic domain3,4.
The optimal design of siRNAs to achieve appropriate guide strand selection and efficient target gene silencing is a problem that is largely solved5,6. As such, the application of RNAi in therapeutically relevant contexts is now the subject of intense investigation, with the first clinical trials directed at age-related macular degeneration via local ocular siRNA delivery and at the lung for respiratory syncitial viral infection7,8.
Targeting neurodegenerative disease: key challenges
With the impressive advances in basic RNA biology and in the clinical applications of RNAi, a question now raised: whether therapeutic RNAi approaches can be made to work effectively in the most demanding cellular environment and within the most complex organ system in order to treat human neurological disease?
There are powerful reasons to pursue this approach, not least the rapidly increasing incidence of neurodegenerative disorders in the aging populations of developed nations and the lack of effective medicines for these diseases. However, the broad range of neurodegenerative disorders, along with the less than adequate understanding of disease pathogenesis in many cases, the fact that whilst many have a defined genetic basis most neurodegenerative disease is sporadic in origin and the complexity of the neurodegenerative process within various nervous system regions, all create significant obstacles to developing effective RNAi therapies. Which neurodegenerative disorders might therefore be best suited to demonstrating proof-of-principle for an RNAi therapeutic agent? Those in which the molecular pathogenesis is well characterised, which have a clear genetic basis involving dominant gain of function, which have localised nervous system pathology and which offer the possibility for localised RNAi delivery, should be at the top of the list (see Figure 1). The prime candidates are most likely to include: Parkinson’s disease (PD), where several dominant genetic forms are now well characterised, and where the neuropathology is focused on dopaminergic neurodegeneration within the midbrain9; Huntington’s disease (HD), a purely genetic disorder comprising trinucleotide repeat expansion gain of function mutations within the huntingtin gene; and the spinocerebellar ataxias (SCA), particularly those which are purely genetic, involve trinucleotide repeat / polyQ expansion mutations as is the case for HD10,11. SCAs result from cerebellar neurodegeneration, but SCA7, for example, also involves retinal degeneration and may be a very attractive candidate for RNAi proof-of-principle clinical studies. The key question is therefore threefold: can neurodegenerative mutant target genes be effectively targeted with RNAi? For example, discriminated from wild-type; can RNAi effector molecules be efficiently delivered to the brain in vivo in order to evaluate proof-of-concept in patients? Will silencing these target genes lead to delayed disease progression or disease reversal?
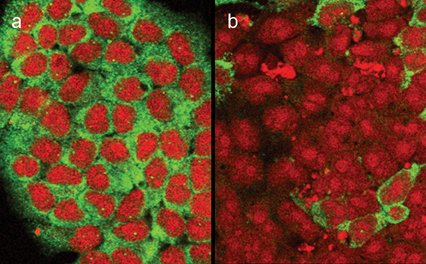

Figure 1: RNAi silencing of a neurodegeneration disease gene (in green) in nervous system stem cells in b, with control treated cells in a.
Silencing neurodegenerative disease alleles
It is known that placing mismatches or mutations within double-stranded siRNAs can attenuate or abolish their ability to silence their mRNA targets effectively12. Thus siRNAs can in principle be utilised for the targeted silencing of transcripts of disease-associated alleles that differ from the wild-type by a single nucleotide. Therefore, can siRNAs be found to discriminate effectively between silencing wild-type and mutant disease genes underlying neurodegenerative disease? Most studies to date have focused on silencing dominant disease alleles that differ from the wild-type by a single nucleotide. However in the case of trinucleotide repeat mutations the targeting of linked single nucleotide polymorphisms (SNPs) has proved an effective strategy. In several of the earliest published studies, centrally placed single nucleotide mismatches between the siRNA guide strand and the wild-type transcript lead to successful allele-specific discrimination.
Miller et al. effectively targeted the mutant Machado-Joseph disease / SCA3 allele through a linked SNP and also effectively silenced a missense Tau mutation using siRNA duplexes and short hairpin RNA (shRNA) approaches13. In other work from the same group the TOR1A gene, causing inherited dystonia, was also successfully targeted14. A third study from our own group demonstrated effective allele-specific silencing of mutations in the acetylcholine receptor, which underlie kinetic disorders of neuromuscular transmission, such as slow channel congenital myasthenic syndrome, using both siRNA and shRNA strategies15.
A definitive study by Schwarz and colleagues attempted to identify the biochemical factors and design variables underlying effective single nucleotide discrimination by siRNAs, such that rational design of optimised discriminatory siRNA sequences might be possible16. This work targeted shortened versions of the mutant SOD1 gene (underlying hereditary amyotrophic lateral sclerosis) and mutant huntingtin gene (underlying HD) in luciferase reporter assays. The authors identified positions within the siRNA guide strand that were most sensitive to mismatches, notably nucleotides in the 3’ region especially the nucleotide at position 16 and demonstrated that purine:purine mismatches confer greater discriminatory efficacy than other types of nucleotide mismatches. In addition, the authors reported that G:U wobbles or single nucleotide mismatches within the ‘seed’ region, a domain essential for target mRNA binding, yielded reduced selectivity compared with mismatches 3′ to this region, such as positions 10 and 16. While this study provides the most detailed analysis to date of the factors governing single nucleotide siRNA discrimination, it is not yet clear to what extent this knowledge can be generalised to all gene targets relevant to neurodegenerative disease. More importantly, given that this was an siRNA in vitro study targeting short transcripts linked to luciferase reporters, it remains to be demonstrated whether such design rules hold true for shRNA-derived dsRNAs, and for the targeting of full-length disease transcripts, ultimately in in vivo disease models.
Even though the mechanistic findings of Schwarz et al.16 have not yet been tested thoroughly in in vitro or in vivo cellular or animal models of neurodegenerative disease, and thus the allele-specific discrimination reported in some studies may be sub-optimal, encouraging data from several studies targeting mutant genes in PD, HD and the SCAs is beginning to emerge. In PD, one of the first genes to be identified in which dominant gain of function missense mutations leads directly to early onset neurodegeneration is the alpha synuclein gene. Targeting the A53T mutation in alpha synuclein, Sapru and colleagues evaluated siRNAs in which the A53T-specific mismatches were placed at positions 9, 10 or 11, and found that effective silencing of the A53T mutant was achieved with the mismatched nucleotide at position 9, with minimal silencing of the wild-type17.
In unpublished studies from our own laboratory we have targeted the alpha synuclein A30P mutation using expressed shRNAs and have attempted to validate some of the findings of Schwarz et al. (Sibley at al. unpublished data). The A30P mutation arises from a G to C substitution in the alpha synuclein gene, creating a strong G:G purine:purine nucleotide mismatch with the wild-type sequence. Luciferase screening of a tiled series of shRNAs across this mutation revealed highly effective allele-specific discrimination with the mismatched nucleotides at positions 12, 14, 15 and most effectively at position 16, corroborating the biochemical significance of this nucleotide position as a 3’ site of consistently low tolerance to mismatches or mutations.
Some of the greatest promise to date has arisen from studies in two laboratories related to HD. As indicated earlier, this disease arises from a CAG repeat expansion in exon 1 of huntingtin gene leading to a polyQ expansion and toxic gain of function in the mutant protein. In an important study targeting the mutant human huntingtin gene, Harper and colleagues demonstrated effective silencing of the mutant transcript in vitro and in vivo using AAV vectors to drive shRNA expression from a U6 promoter. Moreover, using an HD mouse model HD-N171-82Q, containing 82 CAG repeats, the authors were able to demonstrate improvements in the cellular and behavioural disease phenotype following mutant gene silencing18. Given that this study targeted the mutant human huntingtin gene in the mouse, what it failed to do was demonstrate that phenotypic correction could occur via allele-specific silencing, an approach that is likely to be important, possibly even essential, in a clinical study. Work to identify disease-associated SNPs or other strategies for allele-specific silencing in HD is in progress. In a more recent study from a second laboratory the authors demonstrate similar findings, in this case using cholesterol-conjugated siRNAs19. In this study an acute onset HD mouse model was developed via AAV expression of a human mutant huntingtin fragment. Despite successfully correcting the mutant phenotype to some degree, again this was not achieved via allele-specific silencing. Moreover, the relevance of such an acute onset mouse model to human HD is not clear. The final disease group in which promising work has been reported is SCA. Many, although by no means all, of the SCAs are dominantly inherited neurodegenerative diseases caused by expanded CAG trinucleotide repeats within the relevant gene, and hence are purely genetic disorders with a similar molecular pathogenesis to HD. Targeting SCA1, caused by an expanded polyQ tract in the ataxin-1 protein, Xia et al. were able to demonstrate in vivo correction of the cellular phenotype and cerebellar morphology in transgenic mice expressing the human disease allele20. While this study did not demonstrate phenotype correction via allele-specific silencing, its importance, as in the two HD studies referred to earlier, lies in its demonstration as proof-of-principle for the RNAi approach as a therapeutic strategy for neurodegenerative disease. In unpublished work from our own laboratory, we have undertaken an extensive study targeting SCA7, a related disorder that arises due to a polyQ expansion in the ataxin-7 gene (Scholefield et al. unpublished data). Targeting a linked SNP to the ataxin-7 mutation which creates a weak G:U nucleotide mismatch, we have carried out an extensive shRNA screen to optimise allele-specific silencing of this mutant transcript and corroborate and extend the findings if Schwarz et al. We find that single nucleotide mismatches placed 3’ in the shRNA confer the greatest selectivity, although in addition we do find that screens against short transcript targets in luciferase assays are not always predictive of mismatch positions that are most effective against the full-length target. Mismatches at position 16 were again found to be highly efficient for allele-specific silencing, even more so in heterozygous assays in which mutant and wild-type transcripts were co-expressed. Finally the position 16 mismatched shRNAs were able to successfully eliminate ataxin-7 polyQ disease aggregates and correct the cellular phenotype in a disease model; a very encouraging result which suggests that this strategy for allele-specific mutant silencing has real promise for the trinucleotide repeat expansion disorders, including HD.
Evidence that specific silencing of disease associated alleles is likely to delay or reverse disease progression comes from the HD and SCA1 studies discussed above18-20. In addition, an important earlier study in a tetracycline-regulated mouse model of HD21 demonstrated reversal of the pathological and behavioural features following repression of mutant transgene expression. Thus, as discussed in this section, RNAi strategies to attenuate or abolish the expression of disease alleles underlying a range of neurodegenerative disorders are, if successful, likely to result in significant clinical benefit to the patients.
Delivery of RNAi to the nervous system
Before evaluating this RNAi approach for neurodegenerative disease in patients, the other principal hurdle to be overcome is the efficient delivery of RNAi trigger molecules to the appropriate target regions of the nervous system. In general, two delivery approaches are being pursued for RNAi therapeutics4. The use of synthetic double-stranded siRNAs is the simplest approach involving the delivery of a biotherapeutic nucleic acid that may or may not have been chemically or covalently modified in various ways. The second approach is the use of viral expression systemsto deliver expressed RNAi triggers22,23, typically using AAV or lentiviral vector delivery systems12,20,22,24.
A non-viral synthetic siRNA approach has distinct advantages as a small molecule biotherapeutic agent in terms of drug development. The burgeoning number of small biotechnology companies focused on therapeutic siRNA technology including Sirna Therapeutics, Alnylam Pharmaceuticals and others, and the now significant interest being taken by major pharmaceutical companies including Merck, Roche and Novartis testify to this fact. The major limitation of this approach to date, especially in the context of neurodegenerative disease, has been efficient delivery.
Few studies have demonstrated effective target gene silencing following single intracerebral siRNA injections. Even studies such as that by DiFiglia and colleagues referred to earlier19, where some efficacy has been shown, in this case with the use of cholesterol-conjugated siRNAs, the levels of target gene silencing are low. The most impressive results to date have been achieved following chronic siRNA delivery with the use of implanted minipumps25,26. The ‘holy grail’ for the nervous system will ultimately be systemic vascular delivery to the brain. In this respect an intriguing study was published in 2007 by Kumar and colleagues in which they report transvascular siRNA delivery to brain and efficacy in a model of viral encephalitis27. In this work, a rabies virus glycoprotein derived 29mer peptide which binds the acetylcholine receptor (expressed by neuronal and endothelial cells), to which was added a 9mer arginine peptide, the resulting peptide complexed with siRNA, was found to significantly enhance brain siRNA delivery following intravenous injection. Whilst this study is promising, the real significance of this work will be able to be better evaluated when the mechanism is more accurately defined and the findings are reproduced in other laboratories. Nevertheless the identification of novel targeting peptides to enable efficient nervous system delivery may prove a fruitful avenue of research. In another recent high profile study, systemic siRNA delivery to liver has been optimised, in this case using cholesterol-conjugated lipophilic siRNAs which exploit the LDL receptor pathway for liver uptake28,29. To what extent this work is of direct clinical relevance is not yet clear, and it is unlikely to have direct applications for brain delivery. Despite this, the development of efficient siRNA delivery to brain, in particular via the vascular route, remains a high research priority.
In the absence of efficacious, reproducible non-viral siRNA delivery to the nervous system, the most effective strategy to date has exploited the use of viral delivery systems, chiefly adeno-associated virus (AAV) and lentivirus vector systems18-20. In particular the studies referred to earlier by Xia et al. and Harper et al. utilising AAV vectors are the best examples of the power of this approach and the high efficiency delivery and shRNA expression that can be achieved using viral systems. The principal caveat with this approach is that shRNAs may result in toxicity via saturation of the miRNA pathway, when expression is at high levels and unregulated20,30. The solution to this problem is to regulate shRNA dose and expression more tightly, ultimately developing vectors that can be switched off, as well as developing expression systems that utilise natural miRNA structures and sequences to facilitate regulated endogenous processing of the transcript.
At present the viral approach is the one most likely to be utilised in the first clinical studies evaluating RNAi therapies for neurodegenerative disease.
Conclusions
Major advances are being made in RNAi silencing of disease alleles for neurodegenerative disease, and in our understanding of how this can be optimised. PD, HD, and SCAs1-7 are disorders in which the neurodegenerative process is relatively localised and the target genes (in some genetic cases of PD) and molecular pathogenesis sufficiently well characterised for these to represent the most straightforward disease targets for proof-of-concept Phase I clinical trials. SCA7 is particularly promising since it offers the prospect of a local delivery ocular approach to evaluate efficacy in the retina. Such studies are likely to initiate within the next few years, and will give us a clearer indication of the therapeutic potential of RNAi as a 21st century medicine for neurodegenerative disease.
References
- Fire, A., Xu, S., Montgomery, M.K., Kostas, S.A., Driver, S.E., and Mello, C.C. (1998). Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806-811.
- Elbashir, S.M., Harborth, J., Lendeckel, W., Yalcin, A., Weber, K., and Tuschl, T. (2001). Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 411, 494-498.
- Hannon, G.J. (2002). RNA interference. Nature 418, 244-251.
- Kim, D.H., and Rossi, J.J. (2007). Strategies for silencing human disease using RNA interference. Nat Rev Genet 8, 173-184.
- Dorsett, Y., and Tuschl, T. (2004). siRNAs: applications in functional genomics and potential as therapeutics. Nat Rev Drug Discov 3, 318-329.
- Pei, Y., and Tuschl, T. (2006). On the art of identifying effective and specific siRNAs. Nat Methods 3, 670-676.
- Bitko, V., Musiyenko, A., Shulyayeva, O., and Barik, S. (2005). Inhibition of respiratory viruses by nasally administered siRNA. Nat Med 11, 50-55.
- McFarland, T.J., Zhang, Y., Appukuttan, B., and Stout, J.T. (2004). Gene therapy for proliferative ocular diseases. Expert Opin Biol Ther 4, 1053-1058.
- Wood-Kaczmar, A., Gandhi, S., and Wood, N.W. (2006). Understanding the molecular causes of Parkinson’s disease. Trends Mol Med 12, 521-528.
- Orr, H.T., and Zoghbi, H.Y. (2007). Trinucleotide repeat disorders. Annu Rev Neurosci 30, 575-621.
- Soong, B.W., and Paulson, H.L. (2007). Spinocerebellar ataxias: an update. Curr Opin Neurol 20, 438-446.
- Amarzguioui, M., Holen, T., Babaie, E., and Prydz, H. (2003). Tolerance for mutations and chemical modifications in a siRNA. Nucleic Acids Res 31, 589-595.
- Miller, V.M., Xia, H., Marrs, G.L., Gouvion, C.M., Lee, G., Davidson, B.L., and Paulson, H.L. (2003). Allele-specific silencing of dominant disease genes. Proc Natl Acad Sci U S A 100, 7195-7200.
- Gonzalez-Alegre, P., Bode, N., Davidson, B.L., and Paulson, H.L. (2005). Silencing primary dystonia: lentiviral-mediated RNA interference therapy for DYT1 dystonia. J Neurosci 25, 10502-10509.
- Abdelgany, A., Wood, M., and Beeson, D. (2003). Allele-specific silencing of a pathogenic mutant acetylcholine receptor subunit by RNA interference. Hum Mol Genet 12, 2637-2644.
- Schwarz, D.S., Ding, H., Kennington, L., Moore, J.T., Schelter, J., Burchard, J., Linsley, P.S., Aronin, N., Xu, Z., and Zamore, P.D. (2006). Designing siRNA that distinguish between genes that differ by a single nucleotide. PLoS Genet 2, e140.
- Sapru, M.K., Yates, J.W., Hogan, S., Jiang, L., Halter, J., and Bohn, M.C. (2006). Silencing of human alpha-synuclein in vitro and in rat brain using lentiviral-mediated RNAi. Exp Neurol 198, 382-390.
- Harper, S.Q., Staber, P.D., He, X., Eliason, S.L., Martins, I.H., Mao, Q., Yang, L., Kotin, R.M., Paulson, H.L., and Davidson, B.L. (2005). RNA interference improves motor and neuropathological abnormalities in a Huntington’s disease mouse model. Proc Natl Acad Sci U S A 102, 5820-5825.
- DiFiglia, M., Sena-Esteves, M., Chase, K., Sapp, E., Pfister, E., Sass, M., Yoder, J., Reeves, P., Pandey, R.K., Rajeev, K.G., et al. (2007). Therapeutic silencing of mutant huntingtin with siRNA attenuates striatal and cortical neuropathology and behavioral deficits. Proc Natl Acad Sci U S A 104, 17204-17209.
- Xia, H., Mao, Q., Eliason, S.L., Harper, S.Q., Martins, I.H., Orr, H.T., Paulson, H.L., Yang, L., Kotin, R.M., and Davidson, B.L. (2004). RNAi suppresses polyglutamine-induced neurodegeneration in a model of spinocerebellar ataxia. Nat Med 10, 816-820.
- Yamamoto, A., Lucas, J.J., and Hen, R. (2000). Reversal of neuropathology and motor dysfunction in a conditional model of Huntington’s disease. Cell 101, 57-66.
- Brummelkamp, T.R., Bernards, R., and Agami, R. (2002). A system for stable expression of short interfering RNAs in mammalian cells. Science 296, 550-553.
- Paddison, P.J., Caudy, A.A., and Hannon, G.J. (2002). Stable suppression of gene expression by RNAi in mammalian cells. Proc Natl Acad Sci U S A 99, 1443-1448.
- Rubinson, D.A., Dillon, C.P., Kwiatkowski, A.V., Sievers, C., Yang, L., Kopinja, J., Rooney, D.L., Zhang, M., Ihrig, M.M., McManus, M.T., et al. (2003). A lentivirus-based system to functionally silence genes in primary mammalian cells, stem cells and transgenic mice by RNA interference. Nat Genet 33, 401-406.
- Thakker, D.R., Natt, F., Husken, D., Maier, R., Muller, M., van der Putten, H., Hoyer, D., and Cryan, J.F. (2004). Neurochemical and behavioral consequences of widespread gene knockdown in the adult mouse brain by using nonviral RNA interference. Proc Natl Acad Sci U S A 101, 17270-17275.
- Thakker, D.R., Natt, F., Husken, D., van der Putten, H., Maier, R., Hoyer, D., and Cryan, J.F. (2005). siRNA-mediated knockdown of the serotonin transporter in the adult mouse brain. Mol Psychiatry 10, 782-789, 714.
- Kumar, P., Wu, H., McBride, J.L., Jung, K.E., Kim, M.H., Davidson, B.L., Lee, S.K., Shankar, P., and Manjunath, N. (2007). Transvascular delivery of small interfering RNA to the central nervous system. Nature 448, 39-43.
- John, M., Constien, R., Akinc, A., Goldberg, M., Moon, Y.A., Spranger, M., Hadwiger, P., Soutschek, J., Vornlocher, H.P., Manoharan, M., et al. (2007). Effective RNAi-mediated gene silencing without interruption of the endogenous microRNA pathway. Nature 449, 745-747.
- Wolfrum, C., Shi, S., Jayaprakash, K.N., Jayaraman, M., Wang, G., Pandey, R.K., Rajeev, K.G., Nakayama, T., Charrise, K., Ndungo, E.M., et al. (2007). Mechanisms and optimization of in vivo delivery of lipophilic siRNAs. Nat Biotechnol 25, 1149-1157.
- Grimm, D., Streetz, K.L., Jopling, C.L., Storm, T.A., Pandey, K., Davis, C.R., Marion, P., Salazar, F., and Kay, M.A. (2006). Fatality in mice due to oversaturation of cellular microRNA/short hairpin RNA pathways. Nature 441, 537-541.
Matthew JA Wood
Department of Physiology, Anatomy and Genetics, University of Oxford
Matthew Wood graduated in Medicine from the University of Cape Town, working in Clinical Neuroscience before gaining a doctorate in Physiological Sciences from the University of Oxford. He is currently University Lecturer, and Fellow and Tutor in Medicine and Physiology at Somerville College, Oxford. Matthew’s research is in field of gene therapy for degenerative disorders of the nervous system and muscle. The main focus is the investigation of novel therapeutic approaches using antisense oligonucleotides for the modification of mRNA splicing in Duchenne muscular dystrophy and using RNA interference gene silencing approaches for neurodegenerative disease.




