GPCR screening and drug discovery: Challenges and latest trends
Posted: 26 April 2012 |
G-protein coupled receptors (GPCRs) are one of the most popular drug targets today. Almost one third of the approved drugs currently available rely on some kind of interaction with these receptors. The annual revenues are around USD 30 billion (109) and the fact that one quarter of the top US selling drugs are GPCR-related puts this drug target under the drug discovery spotlight. Also, GPCRs are one of the largest families in the human genome, with nearly 1,000 sequences identified as likely to be GPCRs. Among these, around 100 sequences have been confirmed as receptors, but have no known ligand or function. These so-called orphan receptors harbour the highest drug discovery potential. Still, even amongst the non-orphan receptors, only a handful are actually targeted by drugs1-3.
GPCR function is associated with cell sensing of external factors including odorants, taste ligands, light, metals, neurotransmitters, biogenic amines, fatty acids, amino acids, peptides, proteins, steroids and other lipids. The immense possibilities of ligands associated with the huge quantity of receptors led to the association of GPCRs with a large number of physiological and pathological conditions which include pain, asthma, cancer, cardiovascular diseases, gastrointestinal diseases, CNS diseases and others3.
The complexity of the GPCR signalling system
GPCR signalling is a highly complex system based on three major elements: i) a GPCR with the ability to couple with a heterotrimeric guanosine-5′-triphosphate (GTP) binding protein (G-protein), ii) a GTP-transferase active G-protein and iii) a second messenger generating enzyme4,5. A general mechanism, whereby the connection of the ligand to the receptor is coupled to the second messenger forming enzyme through the heterotrimeric G-protein, is largely accepted. Ligand binding to the GPCR causes a change in the receptor conformation that in turn binds and activates the G-protein. The active form of the G-protein is then released from the surface of the receptor, dissociating into its α- and β/γ subunits. Both subunits will then activate their specific effectors, causing the release of second messengers. These messengers are recognised by specific proteins (e.g. protein kinases) leading to their activation and triggering the signalling cascade towards a cellular event. The regeneration of the G-protein is carried out by the hydrolysis of the GTP molecule and re-trimerisation of the G-protein to its inactive form (Figure 1)5,6.
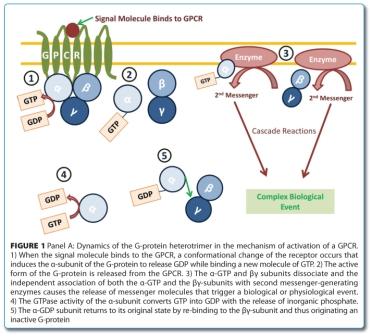

There are two main groups of second messenger releasing enzymes and both are activated or inactivated by different types of G-proteins (Figure 2).
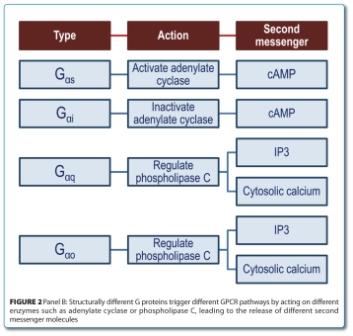

Subtypes Gαs and Gαi either activate or inactivate, respectively, the adenylate cyclase enzyme which converts adenosine triphosphate (ATP) into cyclic adenosine monophosphate (cAMP), releasing inorganic pyrophosphate in the process. Other subtypes such as Gαq or Gαo will alternatively activate the phospho – inositol phospholipase C enzyme (PLC), which hydrolyses phosphatidylinositol-4,5- biphosphate (PIP2) into sn-1,2 diacylglycerol (DAG) and inositol-1,4,5-triphosphate (IP3). The IP3 binds to an endoplasmic reticulum calcium channel, triggering the release of calcium ions into the cytosol. Other IP3 derivatives also have cell function modulation properties. For example, inositol-1,3,4,5-tetraphosphate (IP4) has the ability to facilitate the release of calcium caused by the IP35-7.
GPCR regulation can occur by steric exclusion, receptor internalisation or genetic upregulation/ down-regulation. Two important small families of proteins play a role in these regulation mechanisms: the β-arrestins and the GPCR kinases (GRKs). Like heterotrimeric G-proteins, β-arrestins and GRKs will interact virtually with every GPCR whenever a stimulus is applied. Upon GPCR activation, the second messenger molecules are recognised by GRKs, leading to the phosphorylation of some regions of the GPCR such as the carboxyl terminus. This phosphorylation will not only prevent the association of the GPCR with the G-protein as it will also trigger the recruitment of β-arrestins. These proteins contribute to the regulation by steric exclusion and are also recognised by clathrin coated pit forming elements (e.g. adapter protein 2 (AP2), clathrin and dynamin), leading to the internalisation of the receptor. The β-arrestin molecule is constitutively phosphorylated in the cytosol and is dephosphorylated upon GPCR internalisation. The ubiquitination of the β-arrestin sets both proteins for degradation (Figure 3)5,8-10.
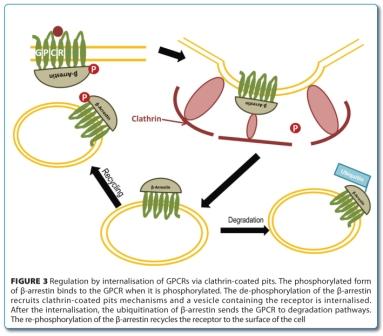

Molecules that interact with GPCRs can be organised in different groups. Considering that, the GPCR system can be defined as a two state system that is either active or inactive. Effectors that interact with the system driving the equilibrium of this to a more active state are called agonists. If the effector interacts with the system by competitive inhibition then it is considered an antagonist. Finally, if the interaction with the GPCR results in further inactivation of the system, then the effector is an inverse agonist11.
GPCR drug discovery
Currently, the main trends in GPCR drug discovery involve not only the search for common ligands such as agonists or antagonists, but also inverse agonists and allosteric modulators (i.e. molecules that interact with the G-protein in places other than the known active site, but that modulate action nevertheless).
Like other receptors, GPCRs are membrane proteins and thus extremely difficult to purify. Consequently, information on 3D structure is only available for a few receptors (e.g. bovine rhodopsin and β2 adrenergic receptor) and approaches for a rational drug design based on receptor structure are difficult to implement. Attempting to predict new candidates from known ligands using computer modelling is also limited by the scarcity of available data12. While some developments in in silico approaches are on-going13, GPCR drug screening is essentially based on testing available molecule libraries. Such libraries can contain millions of different compounds derived either by natural sampling, random synthesis or both, which are tested for their ability to bind or interfere with a given GPCR. Given the large number of molecules in those libraries, the use of High-Throughput Screening (HTS) technologies that enable rapid testing of millions of compounds through miniaturisation and parallelisation is indispensable to perform the screen in a timeframe competitive with market needs14. Ligandbinding assays address the first step in GPCR signalling. Historically, these assays rely on the radioisotopic labelling of compounds. However, the relatively high costs, the limited potential for miniaturisation and hazardousness of the involved protocols have fostered the development of different read-out systems including fluorescence (e.g. time resolved fluorescence-TRF, fluorescence energy transfer-FRET) and luminescence14,15. Typically characterised by a ‘yes or no’ response, ligand-binding assays are useful in determining binding affinities and kinetics but are generally poor in predicting function14.
As drug discovery programmes move towards the development of biologically relevant models, functional assays have been widely adopted. GPCR signalling is a complex mechanism but many of the participants in both action and regulation can be tested to qualify a molecule as a potential GPCR target. The upstream signalling event of GPCR-mediated guanine nucleotide exchange can be monitored by following the accumulation of [35S]GTPγS or europium-labelled GTP. Similar to ligand binding assays, it uses cell membrane preparations. However, the need for a filtration step to separate unbound molecules prevails as a disadvantage for miniaturisation and automation15,16. The exception goes to the scintillation proximity assay bead technology (SPATM) developed by GE Healthcare. Here, membrane preparations are immobilised on SPA beads. Following GPCR activation, membrane accumulation of [35S]GTPγS in the vicinity of the beads produces a SPA signal (www.gelifesciences.com).
The challenge of using cell membrane preparations may be overcome by the use of whole cells that express the GPCR of interest. In this case, monitoring of downstream signalling events is possible. This increases information regarding signalling pathway or modulation activity and provides the tools to perform some molecule / function clustering in early drug discovery stages. Monitoring of downstream signalling events is centered in two main strategies: second messenger accumulation assays and protein redistribution assays. The first strategy comprises the monitoring of the intermediate effectors of a particular signalling pathway. Activation of GPCRs coupled to Gs or Gi is carried out by measuring changes in intracellular cAMP concentrations. Competition assays between endogenous cAMP and exogenously labelled-cAMP, making use of luminescence and fluorescence read-out systems compatible with HTS, are available in the market. The AlphaScreenTM technology from Perkin Elmer is based on competition assays between exogenous, biotin-labelled cAMP and endogenous, non-labelled cAMP. The former is recognised by an anti-cAMP immobilised in acceptor beads. A streptavidin donor bead recognises the biotin-cAMP bead complex. Upon laser excitation, a photosensitiser in the donor beads converts ambient oxygen into a higher energy level state that is translocated to the acceptor bead resulting in light emission20 (www.perkinelmer.com). Competition assays using fluorescent-labelled cAMP with fluorescence polarisation read-out (Perkin Elmer, Molecular Devices, GE Healthcare) or TRF (Cisbio) are also available. The cAMP-Glo kit from Promega uses luminescence derived from dephosphorylation of cAMP dependent protein kinase A (PKA), exogenously added to the system (www.promega.com). Other approaches for cAMP assaying include enzyme fragment complementation (EFC) assays (e.g. Hit HunterTM from DiscoveRx).
GPCR coupling to Gq signalling is usually monitored by following the rise in intracellular calcium resulting from the generation of IP3. Calcium screening can be performed using calcium sensitive fluorescent dyes and fluorometric imaging plate readers (FLIRP). Cell permeable dye formulations are available from several companies including no-wash formulations (e.g. Fluo4 DirectTM from Invitrogen) making these assays simple to execute21,22. Alternatively, cell lines expressing the aequorin calcium sensitive photoprotein are available (e.g. AequoScreen Technology from Perkin Elmer), which provide luminescent readouts23. Competitive binding assays for the detection of IP3 have been developed in particular using fluorescent labelled IP3 in fluorescence polarisation detection schemes (DiscoveRx), SPATM (GE Healthcare) and AlphaScreenTM technology (Perkin Elmer). However, the need for washing procedures or cell lysates is hindering their full integration into HTS-compatible formats15.
Protein redistribution assays focus on the last step in GPCR signalling by following receptor internalisation and translocation across plasma membranes. Since these events are a ubiquitous response to agonistinduced activation, assays targeting receptor internalisation are considered a universal screening platform19. One important aspect is that the majority of these screens rely on the availability of sophisticated cell imaging equipment, generally referred as High Content Screening (HCS) platforms24. Currently, image acquisition has moved from traditional fluorescence optics to confocal-based micro – plate readers with massive improvements on both imaging quality and multiplexing capabilities. A frequent strategy is to use recombinant cell lines expressing a greenfluorescent protein (GFP) tagged-GPCR or an epitope-GPCR construct that is recognised by a tagged antibody. Upon activation, the tagged receptors move towards intracellular endosomes giving rise to defined labelled spots within the cell cytosol. Some drawbacks of this approach are associated with a defective targeting of the fused constructs to the membrane and altered pharmacological response due to the modification of the receptors. The rapid translocation of cytosolic β-arrestins to the plasma membrane following agonistdependent receptor phosphorylation offers a different screening opportunity17,24. The use of cells expressing fused GFP-β-arrestin constructs is the assay principle behind the TransfluorTM technology (www.moleculardevices.com) whilst a non-imaging system based on EFC is the core of the Path HunterTM arrestin technology from DiscoveRx.
The new paradigms driving GPCR drug discovery
Clearly, GPCR drug discovery is now supported by well-established technologies that offer different perspectives of the entire signalling pathway. The choice of the technology to use is not trivial and relies basically on the amount of investment a promoter is willing to support, objectives, resources and team expertise. Generalist approaches are available that are suitable to test a battery of ligands, whose number can be further narrowed-down through subsequent down-streamed functional assays. However, choosing a pathway-biased strategy carries with it the risk of missing ligands that act on different pathways. Moreover, the further we move from the binding event, the higher the risk is of obtaining false positives due to signal amplification. Orphan GPCRs are unlikely to benefit from pathway driven technologies unless some molecule clustering is known in advance14. Universal platforms comprising protein translocation assays appear more appropriate in this context. Interestingly, β-arrestin translocation to membrane receptors, although a late event, is usually associated with a compromise towards receptor internalisation and thus higher signal to noise ratios can be obtained19. Alternatively, cell lines expressing promiscuous G-proteins that couple many receptors to PLC and IP3, can provide for a second type of universal screens25.
The implementation of the technologies described above, alone or in combination, has provided valuable information that is currently impacting drug discovery programmes and opening the way for completely new approaches. One such example is the evidence of GPCR oligomerisation between close or distinct GPCR family members, which may alter ligand binding and G-protein selectivity. Moreover, it is now widely recog nised that GPCR signalling is a product of a complex network, resulting from positive and negative feed-backs from multiple GPCRs in a process designed as receptor cross-talk26. Perhaps one of the most extraordinary findings is related to functional selectivity, a process in which certain ligands stabilise different receptor conformations in such a way that certain pathways are favoured in detriment to27.
All of these findings suggest that the heterogeneous distribution of GPCR population in vivo may result in cell-specific signalling phenotypes26,27. Thus, the general strategy of using reconstituted cell lines over-expressing a particular GPCR is being replaced by more realistic models. Advances in cell culture technology are now providing the means to study primary cells or tissues in HTS or HCS formats28. Cell microarray technology is envisaged to drive multiplexing to unprecedented degrees29. New detection strategies are being developed based on nonlabelled systems such as impedance-based sensors (e.g CellKeyTM) or resonance wave-guide grating (e.g. BINDTM)30. Microfluidic cell technology, although in an early stage, holds the potential for a controlled temporal and spatial resolution thus allowing the interaction with the different ligands to be monitored in time frames equivalent to the in vivo environment31. Microfluidic-based cell assays also offer a promising approach to single cell or even sub-cellular resolution if integrated with miniaturised sensors1.
As GPCR signalling continues to be disclosed, new mechanisms and ligands are revealed and new possibilities are offered by biosensor technologies, one thing is certain: GPCR signalling will continue to challenge and inspire future generations of scientists and engineers.
References
- Prazeres, D.M.F., et al. Towards a high-throughput drug discovery platform for the screening of GPCR targets in cells. in 1st Portuguese Meeting in Bioengineering (ENBENG), 2011
- Schlyer, S. and R. Horuk, I want a new drug: G-proteincoupled receptors in drug development. Drug Discov Today, 2006. 11(11-12): p. 481-93
- Sullivan, F., Emerging technologies in GPCR-based drug target receptors. Frost & Sullivan Annual Report, 2009
- Lefkowitz, R.J., Seven transmembrane receptors: something old, something new. Acta Physiologica, 2007. 190(1): p. 9-19
- Cabrera-Vera, T.M., et al., Insights into G protein structure, function, and regulation. Endocr Rev, 2003. 24(6): p. 765-81
- Hendriks-Balk, M.C., et al., Regulation of G proteincoupled receptor signalling: focus on the cardiovascular system and regulator of G protein signalling proteins. Eur J Pharmacol, 2008. 585(2-3): p. 278-91
- Lagerstrom, M.C. and H.B. Schioth, Structural diversity of G protein-coupled receptors and significance for drug discovery. Nature Reviews Drug Discovery, 2008. 7(4): p. 339-57
- Kristiansen, K., Molecular mechanisms of ligand binding, signalling, and regulation within the superfamily of G-protein-coupled receptors: molecular modeling and mutagenesis approaches to receptor structure and function. Pharmacol Ther, 2004. 103(1): p. 21-80
- Moore, C.A.C., S.K. Milano, and J.L. Benovic, Regulation of receptor trafficking by GRKs and arrestins. Annual Review of Physiology, 2007. 69: p. 451-482
- Shenoy, S.K. and R.J. Lefkowitz, Trafficking patterns of beta-arrestin and G protein-coupled receptors determined by the kinetics of beta-arrestin deubiquitination. Journal of Biological Chemistry, 2003. 278(16): p. 14498-14506
- Brink, C.B., et al., Recent advances in drug action and therapeutics: relevance of novel concepts in G-protein-coupled receptor and signal transduction pharmacology. Br J Clin Pharmacol, 2004. 57(4): p. 373-87
- Jacob, L., et al., Virtual screening of GPCRs: an in silico chemogenomics approach. BMC Bioinformatics, 2008. 9: p. 363
- Schlyer, S. and R. Horuk, I want a new drug: G-proteincoupled receptors in drug development. Drug Discovery Today, 2006. 11(11-12): p. 481-493
- Sewing, A. and D. Cawkill, High-throughput lead finding and optimisation for GPCR targets. GPCRs: From Deorphanization to Lead Structure Identification, 2007. 2: p. 249-267
- Greasley, P.J. and F.P. Jansen, G-protein-coupled receptor screening technologies. Drug Discovery Today: Technologies, 2005. 2(2): p. 163-170
- Heilker, R., et al., G-protein-coupled receptor-focused drug discovery using a target class platform approach. Drug Discovery Today, 2009. 14(5-6): p. 231-240
- Thomsen, W., J. Frazer, and D. Unett, Functional assays for screening GPCR targets. Current Opinion in Biotechnology, 2005. 16(6): p. 655-665
- Wunder, F., et al., Functional cell-based assays in microliter volumes for ultra-high throughput screening. Combinatorial Chemistry & High Throughput Screening, 2008. 11(7): p. 495-504
- Eglen, R.M., Functional G protein-coupled receptor assays for primary and secondary screening. Combinatorial Chemistry & High Throughput Screening, 2005. 8(4): p. 311-318
- Degorce, F., et al., HTRF: A technology tailored for drug discovery – a review of theoretical aspects and recent applications. Curr Chem Genomics, 2009. 3: p. 22-32
- Marshall IC, Owen DE, and M. S., Measuring Ca2+ changes in multiwell format using the fluorometric imaging plate reader. Methods in Molecular Biology, 2006. 312: p. 125-131
- Hodder, P., et al., Miniaturization of intracellular calcium functional assays to 1536-well plate format using a fluorometric imaging plate reader. Journal of Biomolecular Screening, 2004. 9(5): p. 417-426
- Ungrin, M.D., et al., An automated aequorin luminescence-based functional calcium assay for Gprotein- coupled receptors. Analytical Biochemistry, 1999. 272(1): p. 34-42
- Ross, D.A., et al., Multiplexed assays by high-content imaging for assessment of GPCR activity. Journal of Biomolecular Screening, 2008. 13(6): p. 449-455
- Kostenis, E., M. Waelbroeck, and G. Milligan, Techniques: Promiscuous G alpha proteins in basic research and drug discovery. Trends in Pharmacological Sciences, 2005. 26(11): p. 595-602
- Breitwieser, G.E., G Protein–Coupled Receptor Oligomerization. Circulation Research, 2004. 94(1): p. 17-27
- Kenakin, T. and L.J. Miller, Seven transmembrane receptors as shapeshifting proteins: the impact of allosteric modulation and functional selectivity on new drug discovery. Pharmacol Rev, 2010. 62(2): p. 265-304
- Fernandes, T.G., et al., 3D cell culture microarray for high-throughput studies of stem cell fate. Abstracts of Papers of the American Chemical Society, 2011. 241
- Chen, P.-C., Y.-Y. Huang, and J.-L. Juang, MEMS microwell and microcolumn arrays: novel methods for high-throughput cell-based assays. Lab on a Chip, 2011. 11(21): p. 3619-3625
- Fang, Y., A.G. Frutos, and R. Verklereen, Label-free cellbased assays for GPCR screening. Combinatorial Chemistry & High Throughput Screening, 2008. 11(5): p. 357-369
- Wu, M.-H., S.-B. Huang, and G.-B. Lee, Microfluidic cell culture systems for drug research. Lab on a Chip, 2010. 10(8): p. 939-956
About the authors
Sofia M.A. Martins received her PhD degree in Biotechnology from the Technical University of Lisbon (IST), where she worked on the development of particle-based miniaturised formats for DNA detection. Currently, she has a Postdoctoral position at IST and her current research interests are focusing on cell-chips aiming at the characterisation of cell populations and evaluation of cell signalling mechanisms.
João R.C. Trabuco received his BSc in Biochemistry in 2009 from the University of Lisbon (FCUL), and an MsC in Biotechnology in 2011 from the Technical University of Lisbon (IST). Currently he holds a position as a research assistant at IST, working in cell-chip technology development. His main interests are general microscopy, cell imaging techniques, imaging software development and molecular biology.
Gabriel A. Monteiro is a biologist with a PhD in Biotechnology (1998). Currently, he is an assistant professor at the Technical University of Lisbon (IST) and researcher at the Institute for Biotechnology and Bioengineering, Portugal. His main research area is in the development, production and purification of plasmids to DNA vaccination.
Duarte Miguel Prazeres received his PhD in chemical engineering from the Technical University of Lisbon (IST), Lisbon, Portugal, in 1993. He is currently an associate professor in the Department of Bioengineering at IST and a senior researcher in the bioengineering research group. He currently works in the areas of plasmid biopharmaceuticals and nanobiotechnology.




