Live cell high content screening in drug development
Posted: 2 August 2008 | Marc Bickle, Max Planck Institute of Molecular Cell Biology and Genetics, Dresden, Germany | No comments yet
Cell-based assays are essential for drug discovery and development as they increase the quality of lead compounds due to their physiological relevance. Toxicological data can be gathered during the early phases of hit selection and verification, reducing costs and attrition rates during clinical trials.
Cell-based assays are essential for drug discovery and development as they increase the quality of lead compounds due to their physiological relevance. Toxicological data can be gathered during the early phases of hit selection and verification, reducing costs and attrition rates during clinical trials.
Cell-based assays are essential for drug discovery and development as they increase the quality of lead compounds due to their physiological relevance. Toxicological data can be gathered during the early phases of hit selection and verification, reducing costs and attrition rates during clinical trials.
Amongst the various cell-based assays that are currently being used, high content screening (HCS) has enjoyed particular interest and is now widely considered as an essential tool for the drug discovery and development industry and is implemented at every step of the process. This powerful technology is used for target validation1,2, primary screening3, secondary screening4, structure-activity relationship studies5,6 and ADMET (adsorption, distribution, metabolism, excretion and toxicity) studies7. Several factors have contributed to the success of HCS.
Firstly, it is arguably the cell-based assay that can yield the most data due to the richness of information contained in the images. Thus, not only quantitative information such as intensity or number of structures can be obtained, but also descriptive qualifiers, such as size, morphology and texture of objects. Secondly, it is one of the few technologies that can measure spatial data, allowing the assay of translocations events and distribution of objects in regards to each other or to other objects (i.e. vesicle clustering or vesicle distance to plasma membrane). Thirdly, due to the analytical power of modern image analysis, subpopulations can be analysed, making HCS a very sensitive technique, allowing a person to screen for rare events. Lastly, due to the multiplexing capability of HCS, toxicity can be directly assayed by using appropriate markers, instead of being indirectly inferred by reduced cell numbers. Also, several parameters can be screened in parallel, allowing for screening and counter-screening to be performed in a single assay. All these properties facilitate the decision-making process during hit prioritisation and lead development.
In recent years, live cell imaging has emerged as a new trend in HCS and most vendors of HCS readers now offer versions with incubation chambers and environmental control. Live cell imaging adds a further level of complexity to HCS and poses many new technological challenges. This article will review some of the advantages and challenges of kinetic HCS and discuss future directions.
The power of live cell HCS
The vast majority of high content screens are based on end point assays, in which cells are exposed for a predetermined amount of time to perturbing agents such as chemical compounds or RNAi and then fixed. This has the major advantage of uncoupling the production of the cellular assay and the imaging process, facilitating the screening procedure and increasing the throughput. The major disadvantage of end point assays is that the incubation time is the same for all wells, irrespective of the properties of the perturbing agent. In some cases, the incubation time will be insufficient and no phenotype will be scored, whereas for others, incubation time will be too long and secondary effects will be scored. Another drawback of end point assays is that they are more labour intensive than live cell assays, as fixing and staining procedures are time-consuming and laborious. Furthermore, depending on the antibody used, end point assays can be very costly. In live cell imaging, none of the fixing, antibody staining and washing steps are necessitated. Cellular structures are generally tagged with fluorescent proteins and the cells can be visualised with no further manipulations. Sometimes fluorescent dyes are added to the cells, which is not labour intensive, nor are dyes generally costly.
A drawback of performing kinetic HCS is that the throughput is even lower than for end point HCS. As cells have to be either visualised several time of long period of time (for observation of chronic effects) or movies recorded once but for a certain period of time (for recording acute responses), the time spent recording each well is significantly longer than for end point assays. As throughput is a major concern for primary screening, kinetic HCS will only be used for special applications using focused libraries.
Examples of live cell HCS applications
Several assays require the usage of live cell imaging and cannot be carried out as end point assays. For instance, recording calcium waves in neurons require the neurons to be alive. Several such applications have been developed to characterise the properties of compounds affecting various neuronal functions8,9. Measuring electro-chemical potentials across membranes cannot be done reliably in fixed cells and live cell imaging is required. An interesting application of kinetic HCS was reported by O’Brien and colleagues who set up a toxicity assay in HepG2 cells. Markers for plasma membrane potential, mitochondrial membrane potential, calcium concentration and nuclear stains for monitoring chromatin condensation were used to capture several key hallmarks of toxicological events10. Such an assay would not have been feasible with fixed cells and illustrates the power of live cell HCS for compound characterisation. Several groups have also resorted to live cell HCS to monitor rare events such as mitosis11,12. Typically, only a very few number of cells in a population undergo mitosis at any given time. In order to obtain statistically significant data, either a very large population has to be sampled or a small population can be monitored over time, as each cell is bound to undergo mitosis. The latter method has the advantage of avoiding scoring secondary events, as a common phenotype of cells in mitotic arrest, is that to induce apoptosis if arrested at mitotic checkpoints for too long periods of time. Using live cell imaging, the cell cycle of every cell in the field of view can be followed over time, leading to the recording of many mitotic events and classification of the primary phenotype.
Live cell HCS is also required for the characterisation of dynamic events. For instance, the first live cell large-scale screening report was carried out in C. elegans using time lapse differential interference contrast microscopy13. Systematic RNAi of genes found on chromosome III of C. elegans was applied to identify genes involved in early embryogenesis. This screen could only be done in live animals, as errors in the first cellular divisions of the fertilised egg leads to various end phenotypes that do not reveal any information on the original aberrant event. In order to know what went wrong, the early cellular divisions have to be followed live within the uterus of the hermaphrodites.
Another application requiring live cell HCS is the monitoring of subcellular localisation changes. The first high content screen using live cells and automated image analysis and phenotype classification was reported by Conrad and colleagues using arrays of human cells14. Their technology involves the usage of microscope slides carrying spots of gelatin, transfection reagent and cDNA constructs. Seeded cells that adhere to the spots on the microscope slides become transfected. The cells were then imaged on an automatic wide-field microscope and the expression patterns of various GFP constructs were automatically recorded, analysed and classified. By using live cells this approach could potentially record subcellular localisation changes during the cell cycle or due to stimulation with growth factors.


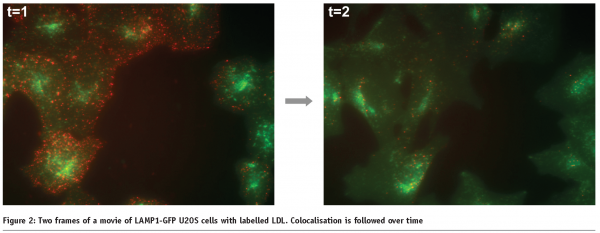
Measuring transport processes, such as vesicular transport or phagosome maturation, also require live cells. We are currently developing a live cell HCS assay for late endosomal transport. In this system, we are tracking individual late endosomes labelled with a LAMP1-GFP construct and assay the speed and movement characteristics after compound addition (Figure 1). In this system we also monitor the increase of colocalisation of vesicles over time after the addition of labelled cargo, as vesicles mature from early endosomes to late endosomes. In this fashion, we assay the effect of compounds on the kinetics colocalisation (see Figure 2).

Migration of cells is important for many processes, ranging from metastasis to wound healing to neuronal development. Migration assays require live cells and such screening assays have been reported in the literature15.
Lastly, a very valuable aspect of live cell HCS for drug development is the pharmacokinetic data that can be obtained from kinetic experiments. As the activity of compounds can be assayed over time, it is possible to determine the kinetic parameter of the mode of action of a compound. It allows us to assess whether a compound is metabolised or stable within cells. Also, compounds do not all act with the same kinetic and it is critical in order to determine the true IC50 to test the activity at different time points. Compounds might not show maximum activity after one hour but will after four hours. In summary, performing kinetic experiments allows determining true IC50 values and obtaining data on the metabolic stability of compounds.
Labelling for live cell HCS
Most organic dyes are prone to photobleaching, limiting their usefulness in live cell imaging. As antibodies cannot be used with live intact cells, labelling techniques are mostly limited to the use of tagged transgenic proteins. Green Fluorescent Protein (GFP) from Aequoeia victoria and its variants have been crucial for the development of live cell imaging, as this family of fluorescent proteins shows little toxicity, is bright, and has excellent photostability properties. This confers to fluorescent proteins superior monitoring properties compared to organic dyes. New fluorescent proteins have been isolated from reef corals17 so that fluorescent proteins with different properties (i.e. absorption/emission maxima, brightness, folding kinetics, oligomerisation) are available16. A new generation of enzymatically activated fluorescence labelling tools has recently emerged such as HaloTag™18 and SnapTag™19. These new tools rely on the enzymatic activity of small proteins fused to the protein of interest. Substrates linked to various fluorescent dyes are added to the cells and become covalently linked to the enzyme, thereby fluorescently labelling the protein under study. The advantage of such systems is that by changing the dye linked to the substrate, different fluorescent absorbtion and emission properties can be obtained with a single transgenic cell line. A further attractive feature of such systems is that they allow carrying out pulse-chase experiments.
All the above technologies involving transgenic construct have the drawback that the proteins of interest need to be expressed at unphysiological levels. It is therefore crucial to determine that the biology under study is not too affected by the experimental conditions.
Many HCS readers have now incorporated brightfield imaging capacities, which has two major advantages. First, morphological information about the cell can be extracted without using a fluorescent channel, so that more information can be gained in a multiplexed experiment. Second, the white light used for brightfield is not very damaging to the cells and the exposure time is very short, so that this imaging mode is relatively benign to the cells. Presently, full advantage of brightfield imaging cannot be extracted, as there are currently few image analysis solutions that can reliably find the edge of cells. We have implemented such an image analysis to segment reliably the cytoplasm of mouse macrophage cell line Raw264.7 (Figure 3). Many companies are currently trying to find solutions, so this problem should be solved in the future. By using plates with glass bottoms instead of plastic, polarised white light imaging should become feasible. Contrast is much more important in differential interference contrast microscopy or Nomarski optics, so that image analysis should be able to extract more information out of the images.


Imaging in kinetic HCS: every photon is sacred!
In order to avoid photodamage and photobleaching, exposure to light has to be kept at a minimum. This constraint impacts on the hardware for recording the images and the image analysis process. In order to reduce light induced damage, the HCS reader should be of the highest quality and have the best available objectives, emission filters and most sensitive cameras available. Regrettably, many HCS readers are equipped with outdated optical technologies, which do not allow for ideal imaging conditions. In order to reduce the light damage, exposure times have to be kept at a minimum, which results in noisy images. Due to the small signal to noise ration of most live cell images, the image analysis process is challenging. This might result in analysis solutions that are computationally more demanding than for higher quality images of end point assays. A further limitation imposed by imaging platforms is the speed of acquisition of the charge coupled device (CCD) camera, the speed of the shutters in the microscope, the speed of integration of the camera and the speed of recording of the computer that will determine the maximum acquisition rate of the microscope. Some biological events are too fast to be recorded reliably with current HCS readers.
Software challenges in live cell image analysis
Very few software available on the market for kinetic HCS offer tracking of objects capabilities. Nearly all of the software of kinetic platforms, segment objects in each time frame and extract features for the objects at each time point. The objects in the individual time frames are not linked to each other and thus no tracking can be performed. Tracking is not a necessity for many kinetic applications. If recording the translocation of a protein to the nucleus over time at different doses of compounds, a statistical analysis of the sub-cellular distribution of the marker in the population of cells is sufficient. The capability to track individual cells and individual responses would add sensitivity to the recording and give some insight into the homogeneity of the cellular response. The cellular response could then also be correlated to other events, such as specific cell cycle events or cell differentiation events. Tracking is therefore not absolutely required for kinetic studies but would refine and enhance the analysis. Algorithms exist to track objects based on probabilistic calculations, which estimate the likelihood that objects found near to each other in different time frames are the same20. To our knowledge, there is only one commercially available software for HCS that exists which has tracking capabilities, which is Kalimoscope (http://www.kalaimoscope.com). Definiens has announced that their latest version that is to be soon commercialised will have tracking capability for 3D and 4D imaging. It is also possible to use MatLab image processing toolbox that contains tools for tracking and some groups have developed their own tracking algorithms to track cells15.
Conclusions
We predict that live cell HCS will become more and more popular in the drug development field in the future. The reasons for this are similar to the reasons which made end point HCS so successful in the first place: a functional read out in a physiologically relevant environment with a wealth of information that allows to better predict the chances of success of lead compounds. In the first instance, kinetic experiments will mainly be carried out in the phases preceding and following primary screening such as target discovery/validation, lead optimisation and ADMET studies. We think that some primary screening campaigns of focused libraries will also be carried out, especially in applications that absolutely require cells to be alive. The instrumentation for live cell imaging should improve, as the optical components of HCS readers improve. Another area where progress is required is image analysis of brightfield images, but solutions should be forthcoming in the future.
References
- C.S. Collins et al: A small interfering RNA screen for modulators of tumor cell motility identifies MAP4K4 as a promigratory kinase. Proceedings of the National Academy of Sciences, 103(10): pp. 3775-3780, 2006.
- A. Friedman and N. Perrimon: A functional RNAi screen for regulators of receptor tyrosine kinase and ERK signalling. Nature, 444(7116): pp. 230-234, 2006.
- G. Xu et al: A high-content chemical screen identifies ellipticine as a modulator of p53 nuclear localization. Apoptosis, 13(3): pp. 413-422, 2008.
- F. Zanella et al: An HTS Approach to Screen for Antagonists of the Nuclear Export Machinery Using High Content Cell-Based Assays. ASSAY and Drug Development Technologies, 5(3): p. 333, 2007.
- D. Haasen et al: Pharmacological Profiling of Chemokine Receptor-Directed Compounds Using High-Content Screening. Journal of Biomolecular Screening, 13(1): pp. 40-53, 2008.
- K.A. Giuliano et al: Systems Cell Biology Knowledge Created from High Content Screening. ASSAY and Drug Development Technologies, 3(5): pp. 501-514, 2005.
- Chad L. Stoner, E.G.C.S.C.S.L.J.B.J.V.N.V.P.N.S., Implementation of an ADME enabling selection and visualization tool for drug discovery. Journal of Pharmaceutical Sciences, 93(5): pp. 1131-1141, 2004.
- G.K.Y. Chan et al: High Content Kinetic Assays of Neuronal Signaling Implemented on BDTM Pathway HT. ASSAY and Drug Development Technologies, 3(6): p. 623, 2005.
- G.R. Richards et al: A Morphology- and Kinetics-Based Cascade for Human Neural Cell High Content Screening. ASSAY and Drug Development Technologies, 4(2): pp. 143-152, 2006.
- P. O’Brien et al: High concordance of drug-induced human hepatotoxicity with in vitro cytotoxicity measured in a novel cell-based model using high content screening. Archives of Toxicology, 80(9): pp. 580-604, 2006.
- B. Neumann et al: High-throughput RNAi screening by time-lapse imaging of live human cells. Nat Meth, 3(5): p. 385, 2006.
- R.O. Burney et al: A Transgenic Mouse Model for High Content, Cell Cycle Phenotype Screening in Live Primary Cells. Cell Cycle, 15(18): pp. 2276-2283, 2007.
- P. Gonczy et al: Functional genomic analysis of cell division in C. elegans using RNAi of genes on chromosome III. Nature, 408(6810): pp. 331-336, 2000.
- C. Conrad et al: Automatic Identification of Subcellular Phenotypes on Human Cell Arrays. Genome Research, 14(6): p. 1130, 2004.
- A. Bahnson et al: Automated measurement of cell motility and proliferation. BMC Cell Biology, 6(1): p. 19, 2005.
- N.C. Shaner et al: A guide to choosing fluorescent proteins. Nat Meth, 2(12): pp. 905-909, 2005.
- M.V. Matz et al: Fluorescent proteins from nonbioluminescent Anthozoa species. Nat Biotech, 17(10): pp. 969-973, 1999.
- G.V. Los and K. Wood: The HaloTag™, in High Content Screening. pp. 195-208, 2006.
- A. Keppler et al: Labeling of fusion proteins with synthetic fluorophores in live cells. Proceedings of the National Academy of Sciences, 101(27): pp. 9955-9959, 2004.
- Y. Kalaidzidis: Intracellular objects tracking. European Journal of Cell Biology, 86(9): p. 569, 2007.



