Academic contribution to high-content screening for functional and chemical genomics
Posted: 21 July 2007 | | No comments yet
High-content screening (HCS) is defined as multiplexed functional screening based on imaging multiple markers (e.g. nuclei, mitochondria etc.) in the physiologic context of intact cells by extraction of multicolour fluorescence information1. It is based on a combination of advanced fluorescence-based reagents, modern liquid handling devices, automated imaging systems and data processing, as well as sophisticated image analysis software.
Initially, HCS was mainly used by pharmaceutical and biotech companies to discover new therapeutic targets and characterize new chemical leads against those targets. As this technology provides not only morphological, phenotypic and genotypic information, but functional data, it has been established as a powerful tool in modern research and drug development. Today, HCS has the potential to be used in a wide range of applications, such as target identification and validation, primary and secondary screening, mode-of-action studies, the hit-to-lead process, the identification of biomarkers, the exploitation of cytotoxicity and genotoxicity, and the tracking of cellular processes applying living cells to support basic research as well as pharmaceutical R&D. While the primary HCS literature still remains rare, a considerable number of review articles summarizing strategies, progress and developments in HCS have been already published, e.g.2-7. Within this review we are going to be focusing on HCS screens published by academic groups; moreover we would like to give a short overview of new technology developments driven by academic groups.
Technology development by academic groups
Considering the wide range of multidisciplinary components and the huge financial investments required for HCS, one can understand that this technology could initially be used by large pharmaceutical companies only. Although the number of HCS units implemented in academic institutions is continuously growing, there is still a significant gap in infrastructure and technical equipment. Currently, there are more than 30 academic centers in the area of small molecule screening, therefore the Molecular Library Screening Centers Network (MLSCN) – a NIH Roadmap initiative, presents the largest academic consortium8,9, but academic groups will hardly be able to compete with the pharmaceutical industry in terms of throughput and speed. In any case, academic researchers are mainly interested in identifying molecular tools in order to investigate biological processes; they are focused on the study of gene functions and the identification of small molecules as well as their mechanisms of action that specifically perturb protein functions. Nevertheless, pharmaceutical screening relies heavily on academic discoveries in disease-relevant biological processes and in detail characterized targets, therefore leaving an important role for target selection with academia. Academics are also willing to enter long-term, riskier projects. Major contributions of academic groups to the field of high-content screening include; the development of innovative, more complex, demanding and physiologically relevant assays such as applying primary cells10, neurons;11, phagosomes or differentiation models12, and development of new technology approaches, e.g. new RNAi libraries13, different delivery methods14, innovative and/or open-source image analysis that increase the number of parameters dramatically to successfully analyse the increasing complexity of multiparametric cellular assays.
Catalogues of subcellular phenotypes have been described for automated recognition by a machine-learning-based classification method15. Ann Carpenter and co-workers offer the open-source image analysis platform, CellProfiler (www.cellprofiler.org)16,17. Rink et al. developed an experimental strategy to track early endosomes over long periods of time by combining high-speed 4D imaging with novel tracking algorithms18. The accuracy of the tracking algorithm was ascertained by systematically comparing computer-generated and manually followed endosome tracks (the program correctly tracked a structure between successive frames with a probability of 99.4%). Kinetic HCS along with correlating instrument and image analysis development has been described for reporting on chromosome segregation and structure by time-lapse microscopy over several consecutive days19.
Chemical genomics
HCS allows the systematic investigation of large libraries and, depending on the type of library used, it can be divided into two main application areas: functional genomics (genomic libraries, RNA interference) and chemical genomics (large compound libraries). In this small review we will focus entirely on screens in cultured mammalian cells rather than in model organisms.
The most commonly used and most published of all HCS strategies have been translocation assays; based on the discrimination between the cell membrane, the nucleus and the cytoplasm, translocation of fluorescently labelled cellular proteins between the distinct compartments is monitored. In 2004 this approach was used to identify NFAT inhibitors (nuclear factor of activated T cells) in nonlymphoid cells with the potential of targeting calcium mobilization in immunosuppression20. Small organic molecules (Diverse E set, ChemBridge Corp., San Diego) were screened for their ability to inhibit the calcium-triggered nuclear import of GFP-tagged NFATc3 in HeLa cells. Fluorescence images of GFP-NFAT, actin visualized by phalloidin staining and nuclei stained with Hoechst 33258 were acquired with an automated image acquisition system. Out of 74 compounds scoring positive in the primary screen, robust inhibition of NFAT nuclear import could be confirmed in 14 (very diverse in structure) compounds .
Proteins involved in signal transduction pathways take the lead in cancer research, the PI3K/PTEN/Akt signal transduction pathway plays a key role in many tumours. A cell-based, visual and chemical genetic screen for small molecule inhibitors of nuclear export of the transcription factor FOXO1a was carried out in cancer-derived cells lacking functional PTEN protein21. The 18,000 compounds from the NCI Structural Diversity Set, ChemBridge DiverSetE, and a small collection of NCI marine extracts were tested for their ability to re-localize FOXO1a to the nucleus in PTEN null cells. The compounds detected in the primary screen were retested in secondary assays, and structure-function relationships were identified. Nineteen of the compounds that promote retention of FOXO1a in the nucleus are novel general protein export inhibitors. They block the nuclear export of RevGFP and FOXO1a by targeting CRM1, and not by inhibition of other FOXO1a factors in the nuclear transport machinery.
The proteasome complex is a new target in cancer drug development. A stably transfected human embryo kidney cell line, HEK 293 ZsGreen Proteasome Sensor, expressing the ZsProSensor-1 fusion protein was used for the screening of proteosome inhibitors, thereby inhibition led to accumulation of the green fluorescent protein ZsGreen22. When screening part of the LOPAC1280 substance library (Sigma Aldrich), several compounds with effects upon the proteasome were found. After exclusion of artefacts and retesting, eight of the compounds were active; among the hits were disulfiram and ammonium pyrrolidinedithiocarbamate (PDTC).
Morphology-based screening of diverse cell types (four cancer cell lines and one non-cancerous primary cell type) has provided a method for discovering new small-molecules as anti-cancer drug candidates23. Cells were treated with various concentrations of 107 small molecules (comprising of four different protein kinase-inhibitor scaffold) and stained for visualization of the nuclei, Golgi apparatus and microtubules. Cellular phenotypes were quantified by multiple statistical analysis of the morphology, staining intensity and spatial attributes of those cell organelles. Principal component analysis was used to identify four distinct compound-induced phenotypes. They identified hydroxyl-PP as targeting carbonyl reductase 1 (CBR1).
Several publications focus on cell cycle analysis; the power of such an approach was presented by Perlman et al.24. A fully automated image-based centrosome-duplication assay was developed and validated with 480 known bioactive compounds. Subsequently, 35 compounds were identified out of a library of 16,320 small molecules (Diverse E set, ChemBridge) using anti-γ-tubulin staining, therefore five specific centrosome-duplication inhibitors could be confirmed.
A small library of 50 synthesized compounds was subjected to two phenotypic cell-based screens monitoring their influence on cell cycle progression through mitosis and viral entry into host cells25. New modulators of cell cycle progression were found and a total of eight compounds reduced virus infectivity and the relative cell number at least 3-fold. Notably, four of these compounds reduced virus infectivity but did not lower the relative cell number below the threshold. An increase in virus infectivity was not observed.
The effects of 880 small-molecule bioactives or drugs were analysed on the insulin-induced adipogenesis of 3T3-L1 fibroblasts, a cell culture model of fat cell differentiation26. Adipose oil droplets were stained with Oil-Red O ten days after the chemical treatment, and the cells were examined under the microscope. Forty-seven compounds stimulated adipogenesis while thirty-nine compounds blocked differentiation without detectable cytotoxicity.
Microscopic platforms allow the assessment of phenotypic changes that are otherwise difficult to quantify, such as cell migration, wound healing and neurite outgrowth. Yarrow et al. discovered an inhibitor of Rho-kinase through an image-based, high-throughput screen of cell monolayer wound healing27. Epithelial cells (BS-C-1) were seeded in 384-well plates and scratch wounded immediately before the addition of compounds (Chembridge DiverSet E). The nuclei and the actin cytoskeleton were visualized and images were acquired using an automated microscope. The compounds affecting wound healing were further investigated with seven secondary assays.
Functional genomics
The second field of HCS are gain-of-function or loss-of-function screening by cDNA transfer or various techniques of triggering RNA interference, respectively. The majority of siRNA screens in mammalian cells are still dealing with reporter gene or other homogeneous assays, nevertheless, multiparametric assays applying automated microscopy gain importance.
Among the first screens in mammalian systems a set of over 5,000 pools of endoribonuclease-prepared siRNAs (esiRNAs) were transfected into HeLa cells and analysed by a homogeneous proliferation assay based on water-soluble tetrazolium (WST-1) still, but all 275 ‘toxic’ hits were analysed applying time-lapse video-microscopy to observe cell proliferation over 3 days. 37 knock-downs exhibited cell division phenotypes28. The first HCS applying a (kinome-wide) chemically synthesised siRNA library demonstrated an impressive complex network of positive and negative regulation of two distinct endocytosis pathways, those are hijacked also by viruses to infect host cells29.
A focus of siRNA screening is to identify new targets or target combinations for cancer therapy, or sensitizers to existing therapeutic small molecules. Among HCS a new lentiviral vector based library consisting of about 5,000 shRNAs targeting 1,000 genes was applied to a three-coloured mitotic index assay staining for DNA, phosphorylated histone H3 and actin to detect cytoplasmic size and shape13. The library included targeting nearly 500 kinases and 180 phosphatases; primary targets in today’s drug discovery. Similarly to the above mentioned chemical wound healing screen a library of 11,000 individual siRNAs targeting 5,234 human genes was probed for genes promoting tumour cell migration30. 523 siRNAs scored, but interestingly, only in 22 genes did both siRNAs per target exhibit the identical phenotype. For all other positive siRNAs, the second molecule targeting the identical gene remained inactive. This provides evidence for the limitations of the current design of siRNAs and the siRNA technology itself. This clearly calls for screening with more siRNAs per target, and identifying a number of non-overlapping siRNAs to verify and validate true hits (refer also to ref. 7).
A second area of interest is neurobiology, new regulators of neurite outgrowth were determined in a gain-of-function screen by transfecting 105 GFP-tagged human full-length cDNA clones into an engineered rat PC12 subline31. Following the GFP-tagged proteins allowed analysis beyond the pure functional aspect of neurite initiation and elongation. A very effective paper was published recently regarding the discovery of genes required for synapse formation and maintenance32. A valuable screening and hit verification strategy was followed. After transcriptional profiling for up-or down-regulated genes during synapse development, about 160 genes were selected to produce an esiRNA library. Pools targeting individual/two or three different targets simultaneously were transfected into cultured hippocampal neurons and a three-coloured assay was performed at two independent runs at least. The very few reproducible hits were verified by an independent RNAi technology. Three shRNAs per target were pooled and transfected, but subsequently the pools were also deconvoluted. In a final genetic rescue experiment it was demonstrated that the phenotype induced by the knockdown of cadherin-13 could be rescued by introducing simultaneously a cadherin-13 expression plasmid resistant to the shRNAs; showing that the decrease in synapse density is due to specific knockdown of cadherin-13.
Human mesenchymal stem cells (hMSCs) were applied to a large-scale siRNA screen consisting of 10,000 siRNAs targeting 5,000 genes to identify suppressors of osteogenic differentiation after 7 days12 by staining after 7 days for ALP activity, an early marker for osteogenic differentiation. 55 hits were derived, 12 of them further characterized. Unfortunately, the work and paper exhibited some weaknesses and limitations for those interested in the screening details, so the biased image acquisition (obviously no automated image analysis was applied), lack of information on the library, whether the siRNAs were pooled or not, which imaging system was used, how the phenotype was quantified, etc.
Merging chemical and functional genomics
Thus far, only one publication has presented a promising approach to screening about 50,000 chemical compounds in parallel to a genome-wide RNA interference33. 214 genes and 50 small molecule inhibitors of cytokinesis were identified. Analysis of the complex phenotypes showed evidence for a small molecule to specifically inhibit the aurora B pathway. Parallel small molecule and RNAi high-content screening might offer a valuable opportunity to compare interesting chemical hits with the catalogue of RNAi phenotypes to allow target prediction. Clearly, the more parameters a complex assay provides the more accurate the target prediction will be.
Future outlook
A number of significant papers have been published in this young discipline, demonstrating the significant contributions made by academic labs in high-content assay and technology development as well as in high-content screening. We are looking forward to seeing more groundbreaking publications being released as many productive academic screening labs have been established recently.
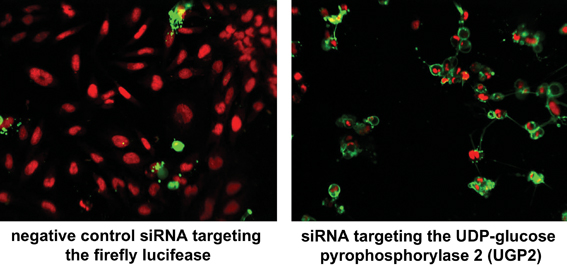

Fig. 1 A high-content siRNA screen was performed to analyse which genes are essential to keep unaltered cell viability. 48 or 96 hours after siRNA transfection, cells were stained with Draq5 and Annexin V without prior fixation. Nuclei appear in red, cells that expose phosphatidylserine (PS) to the outside cell surface are marked in green. The depletion of the UDP-glucose pyrophosphorylase 2 by a targeting siRNA on the right exhibited strong apoptotic phenotypes compared to the negative control treated cells on the left. Multiparametric analysis can be performed by automated image analysis to classify and quantify the various nuclear morphology changes and to measure the presentation of PS on the outside membrane surface. The data contributed to a publication29.
References
- Giuliano KA, DeBiasio RL, Dunlay RT, Gough A, Volosky JM, Zock J, Pavlakis GN and Taylor DL: High Content Screening: A new approach to easing key bottlenecks in the drug discovery process. J. Biomol. Screen. 1997, 2:249–259.
- Mitchison TJ: Small-molecule screening and profiling by using automated microscopy. Chembiochem. 2005, 6(1):33-9.
- Lang P, Yeow K, Nichols A, Scheer A: Cellular imaging in drug discovery. Nat Rev Drug Discov. 2006, 5(4):343-56.
- Haney SA, LaPan P, Pan J, Zhang J: High-content screening moves to the front of the line. Drug Discov Today. 2006, 11(19-20):889-94.
- Rausch O: High content cellular screening. Curr Opin Chem Biol. 2006, 10(4):316-20.
- Nicholson RL, Welch M, Ladlow M, Spring DR: Small-molecule screening: advances in microarraying and cell-imaging technologies. ACS Chem Biol. 2007, 2(1):24-30.
- Krausz E: High-content siRNA screening. Mol. BioSyst. 2007, 3:232-240.
- Gordon EJ: Small molecule screening it takes a village. ACS Chem. Biol. 2007, 2:9-16.
- Lazo JS, Brady LS, Dingledine R: Building a pharmacological lexicon: small molecule discovery in academia. Mol. Pharmacol. 2007, 72:1-7.
- Evans Cousin MA: Simultaneous monitoring of three key neuronal functions in primary neuronal cultures. J Neurosci Methods. 2007, 160:197-205.
- Steinberg BE, Scott CC, Grinstein S: High-throughput assays of phagocytosis, phagosome maturation, and bacterial invasion. Am J Physiol Cell Physiol. 2007, 292:C945-52.
- Zhao Y, Ding S: A high-throughput siRNA library screen identifies osteogenic suppressors in human mesenchymal stem cells. Proc Natl Acad Sci U S A. 2007, 104:9673-8.
- Moffat J, Grueneberg DA, Yang X, Kim SY, Kloepfer AM, Hinkle G, Piqani B, Eisenhaure TM, Luo B, Grenier JK, Carpenter AE, Foo SY, Stewart SA, Stockwell BR, Hacohen N, Hahn WC, Lander ES, Sabatini DM, Root DE: A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen. Cell. 2006, 124:1283-98.
- Bailey SN, Ali SM, Carpenter AE, Higgins CO, Sabatini DM: Microarrays of lentiviruses for gene function screens in immortalized and primary cells. Nat Methods. 2006, 3:117-22.
- Conrad C, Erfle H, Warnat P, Daigle N, Lorch T, Ellenberg J, Pepperkok R, Eils R: Automatic identification of subcellular phenotypes on human cell arrays. Genome Res. 2004, 14:1130-6.
- Carpenter AE, Jones TR, Lamprecht MR, Clarke C, Kang IH, Friman O, Guertin DA, Chang JH, Lindquist RA, Moffat J, Golland P, Sabatini DM: CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biology. 2006, 7:R100.
- Lamprecht MR, Sabatini DM, Carpenter AE: CellProfiler: free, versatile software for automated biological image analysis. Biotechniques. 2007 42(1):71-75.
- Rink J, Ghigo E, Kalaidzidis Y, Zerial M: Rab Conversion as a Mechanism of Progression from Early to Late Endosomes. Cell. 2005, 122: 735-749.
- Neumann B, Held M, Liebel U, Erfle H, Rogers P, Pepperkok R, Ellenberg J: High-throughput RNAi screening by time-lapse imaging of live human cells. Nat Methods. 2006, 3:385-90.
- Venkatesh N, Feng Y, DeDecker B, Yacono P, Golan D, Mitchison T, McKeon F: Chemical genetics to identify NFAT inhibitors: Potential of targeting calcium mobilization in immunosuppression. Proc Natl Acad Sci U S A. 2004, 101(24):8969-8974.
- Kau TR, Schroeder F, Ramaswamy S, Wojciechowski CL, Zhao JJ, Roberts TM, Clardy, J, Sellers WR, Silver PA: A chemical genetic screen identifies inhibitors of regulated nuclear export of a Forkhead transcription factor in PTEN-deficient tumor cells. Cancer Cell. 2003, 4(6):463-476.
- Rickardson L, Wickström M, Larsson R, Lövborg H: Image-based screening for the identification of novel proteosome inhibitors. J Biomol Screen 2007, 12(2):203-210.
- Tanaka M, Bateman R, Rauh D, Vaisberg E, Ramachandani S, Zhang C, Hansen KC, Burlingame AL, Trautman JK, Shokat KM, Adams CL: An unbiased cell morphology-based screen for new, biologically active small molecules. PLoS Biol. 2005, 3:764-776.
- Perlman ZE, Mitchison TJ, Mayer TU: High-content screening and profiling of drug activity in an automated centrosome-duplication assay. Chembiochem. 2005, 6(1):145-51.
- Lessmann T, Leuenberger MG, Menninger S, Lopez-Canet M, Müller O, Hümmer S, Bormann J, Korn K, Fava E, Zerial M, Mayer TU, Waldmann H: Natural product-derived modulators of cell cycle progression and viral entry by enantioselective oxa Diels-Alder reactions on the solid phase. Chem. Biol. 2007, 14(4): 443–451.
- Kawazoe Y, Tanaka S, Uesugi M: Chemical genetic identification of the histamine H1 receptor as a stimulator of insulin-induced adipogenesis. Chem. Biol. 2004, 11:907-913.
- Yarrow JC, Totsukawa G, Charras GT, Mitchison TJ: Screening of cell migration inhibitors via automated microscopy reveals a Rho-kinase inhibitor. Chem. Biol. 2005, 12:385-395.
- Kittler R, Putz G, Pelletier L, Poser I, Heninger AK, Drechsel D, Fischer S, Konstantinova I, Habermann B, Grabner H, Yaspo ML, Himmelbauer H, Korn B, Neugebauer K, Pisabarro MT, Buchholz F: An endoribonuclease-prepared siRNA screen in human cells identifies genes essential for cell division. Nature. 2004, 432:1036-40.
- Pelkmans L, Fava E, Grabner H, Hannus M, Habermann B, Krausz E, Zerial M: Genome-wide analysis of human kinases in clathrin- and caveolae/raft-mediated endocytosis. Nature. 2005, 436:78-86.
- Collins CS, Hong J, Sapinoso L, Zhou Y, Liu Z, Micklash K, Schultz PG, Hampton GM: A small interfering RNA screen for modulators of tumor cell motility identifies MAP4K4 as a promigratory kinase. Proc Natl Acad Sci U S A. 2006, 103:3775-80.
- Laketa V, Simpson JC, Bechtel S, Wiemann S, Pepperkok R: High-content microscopy identifies new neurite outgrowth regulators. Mol Biol Cell. 2007, 18:242-52.
- Paradis S, Harrar DB, Lin Y, Koon AC, Hauser JL, Griffith EC, Zhu L, Brass LF, Chen C, Greenberg ME: An RNAi-based approach identifies molecules required for glutamatergic and GABAergic synapse development. Neuron 2007, 53:217-32.
- Eggert US, Kiger AA, Richter C, Perlman ZE, Perrimon N, Mitchison TJ, Field CM: Parallel chemical genetic and genome-wide RNAi screens identify cytokinesis inhibitors and targets. PLoS Biol. 2004, 2:e379.
About the authors
Eberhard Krausz
Eberhard Krausz is heading the High-Throughput Technology Development Studio (TDS), the High-Content Screening Labs of the Max Planck Institute of Molecular Cell Biology and Genetics (MPI-CBG) in Dresden, Germany. Meanwhile, the TDS consists of independently operating subunits for Assay Development, Kinetic HCS, Instrumentation & Screening, and Data Handling. He has previously held responsible positions for six years in the biotech industry at Cenix BioScience GmbH (Germany) and Cyclacel Ltd. (Scotland, UK) dealing with RNA interference and drug discovery, respectively. Before joining a medical research centre in Munich running a gene therapy project in restenosis, he earned his Ph.D. at the LMU München studying gene regulation in eye lens development. By training he is a microbiologist.
Kerstin Korn
Kerstin Korn is leading the screening and instrumentation unit of the High-Throughput Technology Development Studio (TDS), the High-Content Screening Labs of the Max Planck Institute of Molecular Cell Biology and Genetics (MPI-CBG) in Dresden, Germany. She joined the TDS in June 2004 and contributed significantly to the development and progress of the TDS. After earning her Ph.D. in Biochemistry in 2000 from the University of Leipzig, Germany, Dr. Korn held project leader positions in the biotech industry.




