Streamlining with automation and robotics
Posted: 28 September 2006 | | No comments yet
Protein crystallography has been embraced by the pharmaceutical industry to accelerate and rationalise the drug development process. In this role, success rates, throughput and turnaround times have become key competitive factors, and nearly every stage in the protein crystallography process has been targeted for automation using robotics and advanced software. However, it remains a challenge to combine available technologies, information infrastructure and work-flow protocols into a high throughput protein crystallography pipeline that takes account of the specific objectives and resources of an organisation. This article reviews process outlines and key decision making criteria to assist with the selection of a high throughput protein crystallography strategy.
Modern health care relies extensively on drugs to control the function of proteins that play key roles in disease. Unfortunately, the discovery of new drugs is a formidable task. The traditional approach uses assay-based high throughput screens (HTS) to discover natural or synthetic compounds that inhibit the protein of interest1. Due to low hit-rates, millions of compounds may require screening to find candidates for further development. In addition to the relatively high cost of maintaining huge compound libraries, assay-based screening may miss low affinity inhibitors that could still be promising leads for optimisation.
Empirical activity-based assays are now being complemented by structural biology approaches. Modern high-field NMR of 15N-labeled protein can detect direct binding of compounds with micromolar to low-millimolar affinities and throughput of several thousand compounds per day is now feasible1. Protein crystallography has contributed to HTS in several ways. Virtual ligand screening (VLS) uses computational methods and the known structure of a protein’s active site to predict the binding affinity of reagents in a digital compound library2. Assay-based experimental screens can then be focused on target-specific compounds with a high predicted binding affinity3 and crystallographic ligand fragment library screening has shown promise4,5. Initially, VLS struggled to meet its promise, but recent successes have led to a more widespread adoption of VLS to complement empirical screening6 and for the design of target-specific compound libraries7. Crystal structures can also be used at an early stage to predict the drugability of the target using an array of bioinformatics tools8.
The most valuable role of protein crystallography is in the optimisation of drug candidates. The detailed structure of a protein-prodrug complex defines the key functional groups that contribute to binding affinity and specificity, as well as sites of the molecule that can be derivatised to enhance binding or to introduce additional desirable properties such as improved drug uptake, stability, or solubility. This typically leads to an iterative process of structure-based drug design, compound synthesis, functional characterisation and structure determination of the protein in complex with the most interesting compounds9. In order to use protein crystallography as a platform for structure-guided drug discovery and optimisation, it is critical to establish a rapid and reliable structure determination process. High throughput concepts and instrumentation can greatly contribute to achieving this need.
Protein purification and crystallisation
The crystallographic structure determination process can be subdivided into a number of discrete, interdependent tasks with very different qualities. In the optimal scenario, the experiments are tightly integrated with a data base management system, which interfaces with external data bases and forms the backbone of a knowledge-based information infrastructure system (Figure 1).
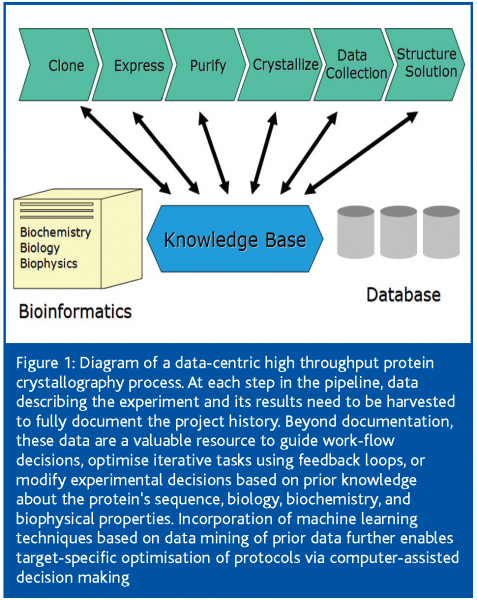

The extensive amount of data generated, especially when successive screening steps are carried out, is frequently underestimated10. Apart from regulatory needs and possible intellectual property aspects, process-information forms the basis for work-flow decisions. While many of these decisions are made by expert users, data stored in a structured machine readable format allow application of machine learning techniques that can drive feedback loops, predict the most promising approach for the next step, or facilitate human decision making. The informatics platform is commonly based on a commercial LIMS package or an application-specific, locally-developed solution centered around a relational database. LIMS packages aimed at the proteomics and crystallography community11,12, are now also being developed by the Structural Genomics centers13.
Protein production
The first major task, protein production and purification, is non-deterministic and for each new target a successful procedure must be established by trial and error. This phase is experiment-driven and robotic automation is increasingly used as a key tool to increase screening success rates and reduce turnaround times and cost.
Based on data from the public structural genomics initiatives, production of a purified, prokaryotic target protein has a success rate of only approximately 30 per cent10. It is clear that for high-value pharmaceutical targets, which often are eukaryotic (human) targets with associated difficulties of expression and presence of post-translational modifications, a significantly greater effort is needed to guarantee a high probability of success. Moreover, the ultimate measure of success is the production of proteins that yield diffraction-quality crystals. Because we cannot predict a priory which approach will be successful, multiple approaches may need to be explored. These alternative approaches can be explored in a parallel or cyclical fashion. The parallel approach uses more resources but has multiple advantages, especially in a pharmaceutical setting where fast turnaround times are critical. Moreover, resource utilisation can be optimised by effective use of automation and efficiencies of scale. It should also be noted that a situation where the parallel approach yields multiple successes is not a waste of resources. Comparison of expression yields enables the selection of the most efficient method, and if different orthologs, mutants or truncation constructs have been produced, all can be carried forward into crystallisation. Finally, when a sufficiently broad protein expression screen fails to yield any success, this can be used to inform important project decisions, leading to either termination or selection of a significantly different approach. Statistically sound early go/no-go decisions are important to manage project resources.
Cost-effective implementation of a parallel protein expression platform requires scalable methods that can be easily automated. This allows for a two-stage approach of low-volume parallel screening followed by scale-up of the most promising conditions14. For cloning, efficient methods are needed to explore the effects of host organism and N- or C-terminal fusion partners. The commercial Gateway (Invitrogen) and In-Fusion systems (Clontech) use enzyme-catalysed homologous recombination to shuttle a single PCR product into multiple expression vectors. Ligation independent cloning (LIC) and classical restriction/ligation methods in combination with a library of compatible expression vectors have been used as alternative, non-proprietary methods. Efficient expression screening of constructs at low volume requires a fast and sensitive assay to quantify expression levels. Anti-His-tag antibodies have been used to probe cell lysates in dot-blot format. The FretWorks assay (Novagen) can quantify proteins containing the 15-residue S-tag in a very sensitive and specific assay that can be readily automated by liquid handling robots (BH, unpublished results).
Protein production can fail at different levels. A common problem in bacterial hosts is the formation of inclusion bodies, especially when expressing eukaryotic genes at high levels. However, numerous examples exist where protein crystals were grown from refolded inclusion body material and incorporation of an inclusion body refolding screen should be considered as part of protein expression screening15. Alternatively, when inclusion bodies are detected, one can screen for conditions that minimise the problem. Reduced expression temperature, weaker or tunable promoters, bacterial strains expressing rare codons or chaperones, or strains with altered di-thiol redox chemistry can all affect protein folding. In this respect it is worth considering whether, during the screening stage, one should aim for maximal expression levels, or for maximal probability to detect expression. Reduced temperatures and/or sub-optimal induction of transcription may actually increase screening success, followed by a quick optimisation of hits before scaling up to production levels.
Protein crystallisation
Protein crystallisation is the least understood aspect of protein crystallography and hundreds of different experimental conditions are typically tested. Crystallisation robots have made it possible to create ever larger crystallisation screens while minimising protein consumption and human labour. However, it has become clear that the most critical factor determining crystallisation success is the protein itself and increasing screen size has diminishing returns16. It therefore makes sense to treat the protein as an important variable and the use of orthologs, truncation analysis and mutagenesis of predicted surface residues has gained a more promising role to address the crystallisation bottleneck. Methylation of lysine residues17 has also been used successfully to give recalcitrant proteins a second chance without having to go back to the cloning stage.
A second area of development is the greater characterisation of the protein sample prior to crystallisation screening. Dynamic light scattering (DLS)18 and protein solubility screening give insight into solution-state aggregation behavior. A new DLS plate reader allows one to optimise monodispersity of the sample in a high throughput format. So far these pre-crystallisation studies have been used to select the protein sample buffer, but not the actual crystallisation screen conditions.
The use of target-optimised protein crystallisation screening has so far been very limited, with the exception of a few screens optimised for special protein classes such as membrane proteins and DNA-binding proteins. Similarly, little progress has been made in using results of past crystallisation screens to guide follow-up screens, apart from optimising hits. Two obstacles need to be overcome to make target-optimised crystallisation screening generally available. First, prior information derived from properties of the target protein, a small protein solubility screen, or a first round crystallisation screen must be transformed into useful knowledge to estimate a crystallisation likelihood function for the target protein16,19. Secondly, target-optimised crystallisation screening requires an efficient technology to create crystallisation cocktails on demand. General purpose liquid handling robots can carry out this task but for routine use this step quickly becomes throughput limiting. A robot that combines screen-creation and crystallisation drop setup has been described20. This robot (Figure 2) uses non-contact dispensing to create crystallisation cocktails in a combinatorial fashion. Throughput in combinatorial dispensing mode is nearly identical to the use of pre-defined crystallisation screens but dynamic screen construction software is needed to exploit the potential of this approach.


For drug development, it is common to seek a structure with a bound ligand or lead compound. In favourable cases, the small molecule can simply be soaked into the crystal. However, when crystals cannot accommodate the ligand, the protein must be co-crystallised together with the ligand. Most ligands are dissolved in DMSO and added in excess, so that after formation of a 1:1 protein-ligand complex the remaining ligand concentration is approximately ten times the binding constant (Kd). This ensures reasonable occupation of the binding site21.
With the increased rate at which crystallisation plates are created, storage, management and evaluation of the plates has become more challenging. Commercially available temperature-controlled incubators with attached automated digital image capture optics solve some of the issues at the hardware level. Solutions that provide crystallisation experiment setup, plate storage and imaging as an integrated package typically include a database driven information structure and user interface. It is important to evaluate the quality and user-friendliness of the software and ensure that data can be exchanged with the existing LIMS that is used for the upstream and downstream stages of the protein crystallisation pipeline. Several products also include automated crystal recognition software that evaluates the images and generates a crystal score. However, it has been difficult to minimise false positives while preventing false negatives. Alternative methods are being developed22,23 but have not yet matured. Protein crystals have been detected using Raman spectroscopy but it is unlikely that this can be applied in a high throughput manner24. In-situ diffraction has also been demonstrated in microfluidic chips25, which could be used for crystal detection. However, it remains to be seen whether these approaches give a lower false negative rate than optical methods, especially when dealing with micro-crystals.
Data collection and structure determination
Once the crystals are safely harvested and stored under cryogenic conditions26 (largely to prevent radiation damage during the subsequent exposure to the intense X-rays), the actual process of the crystallographic structure determination begins, following the steps outlined in Figure 3.
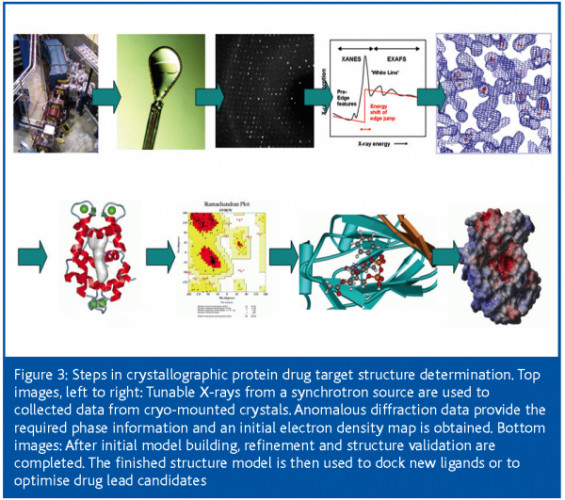

Data collection
Data collection strategies greatly depend on what the ultimate objective of the structure determination seeks to achieve27. Determination of a novel protein structure, especially when experimental phasing is needed, is the most demanding situation and will be discussed first. Given the difficulty and cost of producing protein crystals, it is critical to minimise failure at the X-ray diffraction experiment stage. Failure arises either from damage when the crystal is prepared for data collection, or from an inappropriate data collection strategy. The first step, harvesting the crystal in a cryo-loop28 and cryogenic quenching of the sample, is a manual process and the complex hand-eye coordination is challenging to automate. Although in most environments there is no throughput-driven pressure to automate this step, manual harvesting of crystals from high-density crystal screening plates is difficult and future commercialisation of fully automated or user-guided robotic harvesting aims to increase reliability and reduce crystal loss29.
When available, it is advantageous to mount multiple crystals because crystal quality is the main determinant of data quality and identical-looking crystals can have considerably different diffraction power, as diffraction quality does not necessarily correlate with size or morphological perfection. The concentration and type of cryo-protective additive can also impact diffraction quality and different choices should be explored if possible30. Screening of multiple crystals and cryogenic cooling conditions has been enabled by robots that safely shuttle crystals from a liquid nitrogen storage Dewar to the goniometer and back31. This allows for the rapid collection of test shots from a set of crystals, after which the most promising crystal can be retrieved to collect a full data set. Real-time data analysis is increasingly used to set experimental parameters and select data collection strategies32. Ultimately, it is preferable to solve substructures, calculate phases and build the protein structure while data collection is still ongoing so that the ultimate criterion for data collection success, the ability to solve a crystal structure, can be used to guide the experiment.
New structures that require experimental phasing predominantly use synchrotron radiation combined with anomalous diffraction of seleno-methionine substituted proteins. The recent trend has been towards the use of single-wavelength anomalous diffraction (SAD). SAD phasing requires only one data set compared to two or three for multi-wavelength anomalous diffraction (MAD). A single superior quality data set with less radiation damage is apparently superior to multiple data sets of lower quality. SAD combined with density modification to break the bimodal phase ambiguity yields experimental electron density maps that can be traced automatically. Synchrotron sources are also needed when crystals are too small or diffract too weakly to give adequate data quality at a rotating anode X-ray lab-source. However, new micro-focus rotating anodes and improved X-ray optics have increased the utility of home X-ray sources. This is particularly the case for the screening of many protein-inhibitor complexes where a trade-off in data quality can be tolerated in return for higher throughput and fast turnaround.
Structure determination and analysis
The actual computational task of structure determination is the most mature aspect of the protein crystallography process and, given adequate data quality, phasing, model building and refinement can all be performed with little or no human intervention27. Current efforts are aimed at making the procedure even more robust and less dependent on high resolution data. More work is also needed in the ‘end-game’ where the modeling of alternate side-chain conformations, hydration structure and ligand building is still mostly a manual task.
Significant effort in detailed validation and cleanup of nuisance errors in deposited structure models is necessary to make them suitable for VLS and lead optimisation. Validation of the structure against electron density via real space correlation plots, plausibility based geometry checks, validation of Glutamine, Asparagine and Histidine side chain orientation33, often followed by partial energy minimisation, are applied before the model can be used by docking programs. Similarly, the actual biological quaternary structure34 as well as crystal packing contacts to symmetry related molecules in the crystal structure should be taken into consideration to judge the suitability of a target structure for ligand docking and structure based drug design2.
Conclusion
High throughput protein crystallography has made enormous progress and instruments to automate virtually every step of the process are now commercially available. Although there always remains room for improvement in the hardware, the greater challenge today is to make more efficient use of knowledge. This has been accomplished with great successes in data processing and structure determination but for protein production and crystallisation, brute-force screening is still our main solution to overcome a lack of understanding. Public structural genomics initiatives can play a major role in this area because they have more freedom than drug discovery ventures to share data, and methodological research is often an integral part of their mandate. It is hoped that in the coming years machine learning approaches will give insight into the relative efficiencies of alternate methods, remove screen redundancies and, ultimately, guide the experimental process using empirical knowledge and concurrently collected data while the target moves through the protein crystallography pipeline.
Acknowledgements
BH acknowledges funding by the Alberta Science and Research Authority, the Canadian Foundation for Innovation and the Alberta Synchrotron Institute. BH is an Alberta Heritage Foundation for Medical Research scholars.
References
- Hillisch, A. and R. Hilgenfeld. Modern Methods of Drug Discovery. 2002, Basel: Birkhäuser.
- Abagyan, R.A. and M.M. Totrov. High-throughput docking for lead generation. Curr. Opin. Chem. Biol. 2001 5(4): 375-382.
- Kitchen, D.B., et al. Docking and scoring in virtual screening for drug discovery: methods and applications. Nature Rev. Drug Disc. 2004 3(11): 935-949.
- Blundell, T.L., H. Jhoti, and C. Abell. High-throughput crystallography for lead discovery in drug design. Nature Rev. Drug Disc. 2002 1(1): 45-54.
- Burley, S. The FAST and the curious. Modern Drug Disc 2004 7(5): 53-56.
- Shoichet, B.K. Virtual screening of chemical libraries. Nature 2004 432: 862-865.
- Orry, A., R. Abagyan, and C. Cavasotto. Structure-based development of target-specific compound libraries. Drug Disc. Today 2006 11(5-6): 261-266.
- Ekins, S. Predicting undesirable drug interactions with promiscuous proteins in silico. Drug. Disc. Today 2004 9(6): 276-285.
- Congreve, M., C. Murray, and T. Blundell. Structural biology and drug discovery. Drug Disc. Today 2006 10(13): 895-907.
- Rupp, B. A guide to automation and data handling in protein crystallization, in Crystallization Strategies for Structural Genomics. 2006, International University Line: San Diego.
- Fulton, K.F., et al. CLIMS: Crystallography Laboratory Information Management System. Acta Crystallogr. 2004 D60(9): 1691-1693.
- Haebel, P.W., et al. LISA: an intranet-based flexible database for protein crystallography project management. Acta Crystallogr. 2001 D57(9): 1341-1343.
- Goh, C.S., et al. SPINE 2: a system for collaborative structural proteomics within a federated database framework. Nucleic Acids Res. 2003 31(11): 2833-2838.
- Segelke, B., et al. Laboratory scale structural genomics. J. Struct. Funct. Genomics 2004 5: 147-157.
- Vincentelli, R., et al. High-throughput automated refolding screening of inclusion bodies. Protein Sci. 2004 13(10): 2782-2792.
- Rupp, B. Maximum-likelihood crystallization. J. Struct. Biol. 2003 142(1): 162-169.
- Schubot, F.D. and D.S. Waugh. A pivotal role for reductive methylation in the de novo crystallization of a ternary complex composed of Yersinia pestis virulence factors YopN, SycN and YscB. Acta Crystallogr. 2004 D60(11): 1981-1986.
- Wilson, W.W. Light scattering as a diagnostic for protein crystal growth–A practical approach. J. Struct. Biol. 2003 142(1): 56-65.
- Rupp, B. and J. Wang. Predictive models for protein crystallization. Methods 2004 34(3): 390-407.
- Hazes, B. and L. Price. A nanovolume crystallization robot that creates its crystallization screens on-the-fly. Acta Crystallogr. 2005 D61(8): 1165-1171.
- Danley, D. Crystallization to obtain protein-ligand complexes for structure-aided drug design. Acta Crystallogr. 2006 D62(Pt 6): 569-575.
- Wilson, J. Towards the automated evaluation of crystallization trials. Acta Crystallogr. 2002 D58(11): 1907-1914.
- Bern, M., et al. Automatic classification of protein crystallization images using a curve-tracking algorithm. J. Appl. Crystallogr. 2004 37: 279-287.
- Nagarajan, V. and B. Marquardt. Spectroscopic imaging of protein crystals in crystallization drops. J. Struct. Funct. Genomics 2005 6(2-3): 203-208.
- Doerr, A. Hands-off protein crystallography. Nat. Methods 2006 3(4): 244.
- Garman, E. and C. Nave. Radiation damage to crystalline biological molecules: current view. J. Synchr. Rad. 2002 9: 327-328.
- Rupp, B. High throughput protein crystallography, in Structural Proteomics and High Throughput Structural Biology. 2005, Taylor and Francis: New York. p. 61-104.
- Thorne, R.E., et al. Microfabricated mounts for high-throughput macromolecular cryocrystallography. J. Appl. Crystallogr. 2003 36(6): 1455-1460.
- Rupp, B., et al. Engaging the Final Frontier in Automated High Throughput Crystallography: Robotic Crystal Harvesting. ACA Meeting Series 2006 33: in press.
- Garman, E.F. and S. Doublie. Cryocooling of Macromolecular Crystals: Optimization Methods. Methods Enzymol. 2003 368: 188-216.
- Snell, G., et al. Automatic Sample Mounting and Alignment System for Biological Crystallography at a Synchrotron Source. Structure 2004 12: 1-12.
- Leslie, A., et al. Automation of the collection and processing of X-ray diffraction data – a generic approach. Acta Crystallogr. 2002 D58: 1924-1928.
- Weichenberger, C. and M. Sippl. NQ-Flipper: validation and correction of asparagine/glutamine amide rotamers in protein crystal structures. Bioinformatics 2006 22(11): 1397-1398.
- Henrick, K. and J.M. Thornton. PQS: a protein quaternary structure file server. Trends Biochem. Sci. 1998 23(9): 358-361.
Issue
Related topics
Data Analysis, Informatics, Protein Crystallisation, Protein Crystallography