microRNAs and their potential involvement in HIV infection
Posted: 20 June 2011 |
Treatment and cure of human immunodeficiency virus-1(HIV-1) infection remains one of the greatest therapeutic challenges due to its persistent infection, often leading to acquired immunodeficiency syndrome (AIDS). Although it has been 28 years since the discovery of the virus, the development of an effective vaccine is still far away. Relatively newly discovered microRNAs (miRNA) are a family of small noncoding RNAs that can regulate gene expression primarily by binding to the 3’UTR of targeted transcripts. Understanding how HIV-1 infection affects the host miRNA pathway could shed some new insights related to the basic mechanisms underlying HIV-1 mediated pathologies and T-lymphocyte depletion. Here, we review literature related to the biogenesis of HIV-1 encoded miRNAs, cellular miRNAs that can directly target HIV-1 or essential cellular factors required for HIV-1 replication. We also discuss the feasibility of using miRNAs for HIV-1 therapy.
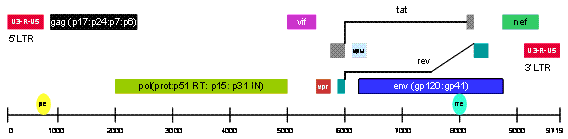

Figure 1 The HIV-1 genome
Treatment and cure of human immunodeficiency virus-1(HIV-1) infection remains one of the greatest therapeutic challenges due to its persistent infection, often leading to acquired immunodeficiency syndrome (AIDS). Although it has been 28 years since the discovery of the virus, the development of an effective vaccine is still far away. Relatively newly discovered microRNAs (miRNA) are a family of small noncoding RNAs that can regulate gene expression primarily by binding to the 3’UTR of targeted transcripts. Understanding how HIV-1 infection affects the host miRNA pathway could shed some new insights related to the basic mechanisms underlying HIV-1 mediated pathologies and T-lymphocyte depletion. Here, we review literature related to the biogenesis of HIV-1 encoded miRNAs, cellular miRNAs that can directly target HIV-1 or essential cellular factors required for HIV-1 replication. We also discuss the feasibility of using miRNAs for HIV-1 therapy.
Human immunodeficiency virus (HIV) -1 is a lentivirus and it is the primary cause of acquired immunodeficiency syndrome (AIDS)1,2. HIV-1 primarily infects CD4+ T memory cells and its infection can be roughly divided into two phases: an acute phase (early phase), which lasts about one to two months after the initial infection, and a chronic infection phase, which can last for upwards of 10 to 20 years before the onset of AIDS3,4. HIV-1 is a relatively small RNA virus, its 9.7 KB RNA genome consists of several secondary structures (LTR, TAR, RRE, PE, SLIP, CRS, INS) and nine genes (gag, pol, env, tat, rev, nef, vif, vpr, vpu) encoding 19 proteins (Figure 1, page 42). There was optimism after the virus was initially discovered to be the cause of AIDS that within a few years a vaccine would be available to both treat and prevent infection. Despite numerous attempts by vaccine experts around the world, highlighted by the failure of the Merck’s STEP trial, HIV-1 vaccine research has been brought back to the starting point. The vaccine used in the STEP trial actually may have promoted, rather than prevented, HIV-1 infection5,6. The failure to create an effective vaccine implies that additional basic research of the HIV-1 infection mechanism is necessary.
microRNAs are 21-23 nucleotide long regulatory, non-coding, small RNAs that repress target gene translation through base pairing to complementary sequences in the 3’ untranslated region (3’UTR) of targeted transcripts7. Studying the interactions of HIV-1 and miRNAs may shed some new insights into the mechanism of host responses to HIV-1 infection. For instance, it is important to understand how HIV-1 infection affects the miRNA pathways and to understand if miRNAs target the HIV-1 genome directly or indirectly to modulate infection. Since microRNAs modulate many different types of viral infection, it was quickly speculated that miRNAs may directly target the HIV-1 viral RNA genome and play a role in modulating HIV-1 replication and infection8. The generation of miRNAs take place through two major steps. First, the production of a precursor hairpin structure (pre-miRNA) from a primary miRNA transcript (pri-miRNA) by the Drosha and DGCR-8 formed microprocessor in the nucleus. Next, the pre-miRNA is exported to the cytoplasm by Exprotin5-RAN-GTP for further processing by Dicer9-11. Dicer excises the pre-miRNA to produce miRNA/miRNA* duplexes (guide strand / passenger strand). Dicer does not act alone, it needs one of the synonymous RNA binding proteins, TRBP and PACT to function as its partner12-14. The ‘guide strand’, is selected and incorporated into RISC, and the other strand, the ‘passenger strand’, is usually degraded15,16. The guide strand will then be loaded onto Argonaute (Ago) to form the RNA induced silencing complex (RISC). The human Argonaute family has four members, Ago1 through Ago4 are closely related and are co-expressed in many cell types. However, endonuclease activity is exclusively associated with Ago217-19. It was found that miRNA mediated repression was mostly dependent on the first eight nts (seed sequence) from the 5’ end of a miRNA and G:U wobble is generally not allowed20. Positions 13 to 16 can help the seed region match and may compensate for weak seed region interactions21. RISC may direct miRNA repressed mRNAs to RNA processing bodies (p-bodies) for deadenylation, decapping and degradation, or alternatively for temporary storage and reuse22-25.
microRNA biogenesis and HIV-1 infection
An important problem is whether or not HIV-1 infection affects individual or global miRNA levels. To affect miRNA or siRNA pathways globally, HIV-1 needs to encode an SRS (suppressor of RNA silencing) protein. It appears that a common phenomenon in viral infections is that virally encoded proteins act as SRSs against the host RNAi pathway components that ordinarily would inhibit their replication26-33. There are other reasons why HIV-1 infection could globally affect miRNA pathways34. Since Dicer and TRBP form the miRNA/miRNA* generation complex, the TAR structure in HIV-1 RNAs could titrate TRBP and affect miRNA biogenesis12,13,35. Other HIV-1 proteins could bind to Dicer or Drosha and affect the miRNA biogenesis pathway. To date there are three publications supporting regulation of the RNAi pathway by the HIV-1 transcription activator Tat36-38. Other studies show that knock down of Dicer or Drosha boosted HIV-1 infection while knock down of TRBP reduced viral infection39. In yet another study, a different conclusion was drawn which suggested that Dicer and its partner TRBP do not play a role in HIV-1 infection via modulation of RNAi. On the other hand, knocking down TRBP or Dicer with siRNAs decreases the expression of viral genes, inhibits viral production and impedes miRNA biogenesis. These data suggest that TRBP contributes mainly to the enhancement of virus production and Dicer does not mediate HIV-1 restriction by RNAi40. A separate study showed that Tat, Tax (HTLV-1), and Tas (PFV-1) failed to inhibit RNA interference in human cells and the stable expression of physiological levels of Tat did not globally inhibit miRNA production or expression in infected human cells41. This conclusion is supported by a recent publication which demonstrated that both HIV-1 Tat and TAR expression do not reduce the efficacy of cellular RNA silencing42. On the other hand, Qian et al. published data supporting Tat as a SRS38 and Nathans et al. reported knock down of Dicer or Drosha boosted HIV-1 infection43. Since Dicer or Drosha knock down also depletes the cellular miRNA population, which in turn affects the normal metabolism of cells, it is not surprising that Dicer or Drosha knock would affect HIV-1 infection. Yeung et al. first reported the RAKE miRNA microarray data of miRNA profiled in pNL4-3 transfected cells versus untransfected cells and concluded that HIV-1 will globally down regulate miRNAs44. Triboulet et al. reported that the polycistronic miRNA cluster miR-17/92 was downregulated in a time course RAKE miRNA microarray in HIV pNL4-3 infected Jurkat cells. This same group showed that Dicer and Drosha inhibited virus replication both in peripheral blood mononuclear cells from HIV-1- infected donors and in latently infected cells. They concluded that the downregulated miR-17/92 cluster would result in upregulation of the target transcriptional co-activator P300/CBP-associated factor (PCAF), a histone acetyltransferase that can interact with Tat and synergise to activate Tat function for efficient viral replication39. It is noteworthy to point out that using the Jurkat cell line to simulate HIV-1 infection may not be totally relevant to an in vivo infection. A study by Chable-Bessia et al. showed that RCK/p54, GW182, LSm-1 and XRN1 negatively regulate HIV-1 gene expression by blocking viral mRNA association with polysomes. Interestingly, their data also showed that knock down of RCK/p54 or DGCR8 resulted in virus reactivation in PBMCs isolated from HAART treated HIV-1 patients45. Houzet et al. reported miRNA profiles in peripheral blood mononuclear cells (PBMC) from 36 HIV-1 seropositive individuals using the RAKE miRNA microarray. Interestingly, they observed downregulation of miR-29a and 29b in both HIV-1 patients and infected PBMCs, and downregulation of miR-29c, miR-26a, and miR-21 in HIV-1 patient blood samples. They proposed specific miRNA signatures can be observed for each of the four classes of HIV-1-positive individuals that were categorised based on their CD4+ T-cell counts and viral loads46. Lastly, Coley et al. showed HIV-1 Vpr can suppress Dicer expression in monocytes47.
Host miRNAs and HIV-1
The role of miRNAs in HIV-1 infection via interactions with cellular miRNA targets is a complicated issue because miRNAs could target both cellular factors that either suppress or activate HIV-1 infection. It is even more complicated when taking HIV-1 encoded miRNAs into consideration. microRNAs involved in HIV-1 infection could be HIV-1 encoded or host encoded and they could also be defined as suppressors or activators of HIV-1 infection. The latter could be further divided as directly targeting HIV-1 transcripts, indirectly targeting HIV-1 via modulation of host factors that are involved in the HIV-1 replication or targeting both the HIV-1 RNA genome and host factors essential for HIV-1 infection.
It is intriguing to determine if host miRNAs play a role in the regulation of HIV-1 gene expression, and if so whether or not they do so by direct targeting of HIV-1 RNA. Under normal conditions, miRNAs function as negative regulators of gene expression by binding to the target message 3’UTR resulting in translational suppression48,49. Because the HIV-1 Nef sequence also serves as the 3’UTR for most of the HIV-1 transcripts, miRNA binding sites in this region may play a critical role in HIV-1 infection50 (Figure 1). Nef has been shown to play a positive role in viral replication and pathogenesis. HIV-1 patients carrying strains with Nef gene deletions have slower progression to AIDS (Reviewed by Foster JL et al.51). Thus, miRNAs targeting the Nef region have the potential to affect HIV-1 pathogenesis. Several host miRNAs that target conserved regions of the HIV-1 genome were predicted in silico52. Huang et al. studied the miRNA expression patterns in activated CD4+ T lymphocytes versus resting CD4+ T lymphocytes and concluded that several host miRNAs (miR-125b, miR-150, miR-28, miR-223, and miR-382), which are highly expressed in resting CD4+ T lymphocytes may target the Nef/3’LTR region and contribute to HIV-1 latency53. They also proposed that the application of the pooled miRNA inhibitors may be used to purge latent state HIV-1 reservoirs54. Similar but somewhat controversial data were reported for monocytes / macrophages by Wang et al.55,56. The determination of the expression levels of those miRNAs enriched in resting CD4 cells relative to activated CD4 cells is difficult to evaluate since the absolute miRNA expression levels in CD4 cells are relatively low according to published large scale miRNA cloning data57,58. It is worth pointing out that highly expressed miRNAs may be important for normal homeostasis of cells. Thus, using pooled antagomirs (anti-micro RNA antisense) to block miRNA function to activate HIV-1 in latent infected cells may be deleterious in uninfected cells. Ahluwalia et al. reported that miR-29a targets the HIV-1 Nef gene and ectopic expression of this host miRNA results in the repression of the Nef protein levels and a reduction of viral levels59. Nathans et al. reported that miR-29a can target HIV-1 and repress its infectivity. They also reported that miR-29a can direct repressed HIV-1 transcripts to P-bodies and this may be a mechanism to keep HIV-1 in a latent state43. Other miR-29 family members can also target the same region52. Interestingly, miR-29 family members are observed to be downregulated in both HIV-1 patients and infected PBMCs46. The mechanism by which HIV-1 infection down-regulates the miR-29 family remains elusive, but Sung et al. reported that host miR-198 restricts HIV-1 replication in monocytes by repressing Cyclin T1 expression60.


Figure 1 The HIV-1 genome
HIV-1 encoded vmiRNAs
In plants, viral dsRNAs are processed to small interfering RNAs (siRNAs) which in turn target the viral genomes for destruction, preventing re-infection by the same virus61. In mammals, a number of virally encoded miRNAs (vmiRNAs) have been described and their functions studied30,32,62-81. An example of a vmiRNA function is that of the Simian virus 40 (SV40) encoded miRNA-1. This vmiRA accumulates late in infections, and is perfectly complementary to early viral mRNAs, therefore, it targets these viral mRNAs for degradation70. It is still debatable as to whether or not HIV-1 encodes vmiRNAs since the data for such virally encoded miRNAs is controversial41,82. HIV-1 encoded candidate small RNAs and their cellular targets were first described from in silico studies83. Subsequently, there have been several papers addressing HIV-1 encoded vmiRNAs. There are three miRBase documented HIV-1 encoded miRNAs: miR-H1- 5p, miR-N367-3p and miR-TAR-5p/3p (Table 1).
Table 1 HIV-1 encoded miRNA/siRNA and their function
| Name | Sequence (miRBase) | References | Function |
| miR-N367 | acugaccuuuggauggugcuucaa | [84, 85] | Suppression of nef |
| miR-H1 | ccagggaggcgugccugggc | [86, 87] | Target human cellular AATF |
| TAR-5p | ucucucugguuagaccagaucuga | [88-91] | Anti apoptosis Chromatin remodeling of the viral LTR |
| TAR-3p | ucucuggcuaacuagggaaccca | [88-91] | |
| vsiRNA1 | uccuuggguucuuaggagc | [36] | Rescuing Env RNA expression |
The controversy in this area is highlighted by a systematic study which showed neither HIV-1 nor human T-cell leukaemia virus type 1 (HTLV-1) expressed significant levels of either siRNAs or miRNAs in persistently infected T cells41. No repeat analyses for miR-N367, miR-H1 and vsiRNA1 have been published. The putative TAR miRNAs are exceptions. These have been reported by two independent groups but the expression levels of the putative TAR-miRNAs are extremely low. Only TAR-3p can be detected by Northern blotting88-91. The cloned sequences from TAR-3p are 17 or 18 nt long, which makes them shorter than the normal 21 to 22 nt lengths of most miRNAs91. Pyro 454 sequencing of HIV-1 derived small RNAs yielded only about 100 reads which mapped to the HIV-1 genome92. Using a Solexa deep sequencing platform, we were able to detect many reads of small RNAs that map to many regions of the HIV-1 genome. We also observed that most of the more abundant deep sequencing reads from HIV-1 do not match the reference sequences in miRBase. While our data showed Tar-3p is present in the small RNA population of HIV-1 infected cells, we detected nearly the same number of reads from the Tar loop region, suggesting that many of the purported HIV-1 encoded vmiRNAs may be RNA degradation products (GS and JJR unpublished data).
In summary, the question of whether or not miRNAs (host or viral) play an essential role in HIV-1 pathogenesis remains unanswered. The regulation of HIV-1 infection by host miRNAs that directly target the viral genome is possible, but it remains uncertain how effective these cellular miRNAs are in influencing HIV-1 infection. The regulation of HIV-1 infection via miRNA targeting of cellular factors is complicated because they affect the expression of genes that could facilitate or inhibit HIV infection. Finally, the question of whether or not HIV-1 infection produces sufficient amounts of HIV derived small RNAs to regulate its infectious cycle remains open.
References
- Chermann, J.C., et al., Isolation of a new retrovirus in a patient at risk for acquired immunodeficiency syndrome. Antibiot Chemother, 1983. 32: p. 48-53
- Barre-Sinoussi, F., et al., Isolation of a T-lymphotropic retrovirus from a patient at risk for acquired immune deficiency syndrome (AIDS). Science, 1983. 220(4599): p. 868-71
- Li, Q., et al., Peak SIV replication in resting memory CD4+ T cells depletes gut lamina propria CD4+ T cells. Nature, 2005. 434(7037): p. 1148-52
- Mattapallil, J.J., et al., Massive infection and loss of memory CD4+ T cells in multiple tissues during acute SIV infection. Nature, 2005. 434(7037): p. 1093-7
- Moore, J.P., et al., AIDS/HIV. A STEP into darkness or light? Science, 2008. 320(5877): p. 753-5
- Oxford, J.S., The end of the beginning: vaccines for the next 25 years. Vaccine, 2008. 26(49): p. 6179-82
- Bartel, D.P., MicroRNAs: genomics, biogenesis, mechanism, and function. Cell, 2004. 116(2): p. 281-97
- Dennis, C., Small RNAs: the genome’s guiding hand? Nature, 2002. 420(6917): p. 732.
- Grishok, A., et al., Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell, 2001. 106(1): p. 23-34
- Hutvagner, G., et al., A cellular function for the RNAinterference enzyme Dicer in the maturation of the let- 7 small temporal RNA. Science, 2001. 293(5531): p. 834-8
- Bernstein, E., et al., Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature, 2001. 409(6818): p. 363-6
- Chendrimada, T.P., et al., TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature, 2005. 436(7051): p. 740-4
- Haase, A.D., et al., TRBP, a regulator of cellular PKR and HIV-1 virus expression, interacts with Dicer and functions in RNA silencing. EMBO Rep, 2005. 6(10): p. 961-7.
- Lee, Y., et al., The role of PACT in the RNA silencing pathway. Embo J, 2006. 25(3): p. 522-32
- Khvorova, A., A. Reynolds and S.D. Jayasena, Functional siRNAs and miRNAs exhibit strand bias. Cell, 2003. 115(2): p. 209-16
- Matranga, C., et al., Passenger-strand cleavage facilitates assembly of siRNA into Ago2-containing RNAi enzyme complexes. Cell, 2005. 123(4): p. 607-20
- Ishizuka, A., M.C. Siomi, and H. Siomi, A Drosophila fragile X protein interacts with components of RNAi and ribosomal proteins. Genes Dev, 2002. 16(19): p. 2497-508.
- Meister, G., et al., Human Argonaute2 mediates RNA cleavage targeted by miRNAs and siRNAs. Mol Cell, 2004. 15(2): p. 185-97
- Okamura, K., et al., Distinct roles for Argonaute proteins in small RNA-directed RNA cleavage pathways. Genes Dev, 2004. 18(14): p. 1655-66
- Doench, J.G. and P.A. Sharp, Specificity of microRNA target selection in translational repression. Genes Dev, 2004. 18(5): p. 504-11
- Lewis, B.P., C.B. Burge, and D.P. Bartel, Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell, 2005. 120(1): p. 15-20
- Wu, L., J. Fan, and J.G. Belasco, MicroRNAs direct rapid deadenylation of mRNA. Proc Natl Acad Sci U S A, 2006. 103(11): p. 4034-9
- Giraldez, A.J., et al., Zebrafish MiR-430 promotes deadenylation and clearance of maternal mRNAs. Science, 2006. 312(5770): p. 75-9
- Behm-Ansmant, I., et al., mRNA degradation by miRNAs and GW182 requires both CCR4:NOT deadenylase and DCP1:DCP2 decapping complexes. Genes Dev, 2006. 20(14): p. 1885-98
- Eulalio, A., et al., Target-specific requirements for enhancers of decapping in miRNA-mediated gene silencing. Genes Dev, 2007. 21(20): p. 2558-70
- Mallory, A.C., et al., A viral suppressor of RNA silencing differentially regulates the accumulation of short interfering RNAs and micro-RNAs in tobacco. Proc Natl Acad Sci U S A, 2002. 99(23): p. 15228-33
- Chapman, E.J., et al., Viral RNA silencing suppressors inhibit the microRNA pathway at an intermediate step. Genes Dev, 2004. 18(10): p. 1179-86
- Calabrese, J.M. and P.A. Sharp, Characterization of the short RNAs bound by the P19 suppressor of RNA silencing in mouse embryonic stem cells. RNA, 2006. 12(12): p. 2092-102
- Zhou, Z., et al., Identification of an RNA-silencing suppressor in the genome of Grapevine virus A. J Gen Virol, 2006. 87(Pt 8): p. 2387-95
- Zhang, X., et al., Cucumber mosaic virus-encoded 2b suppressor inhibits Arabidopsis Argonaute1 cleavage activity to counter plant defense. Genes Dev, 2006. 20(23): p. 3255-68
- Hemmes, H., et al., The NS3 protein of Rice hoja blanca tenuivirus suppresses RNA silencing in plant and insect hosts by efficiently binding both siRNAs and miRNAs. Rna, 2007. 13(7): p. 1079-89
- Haasnoot, J., et al., The Ebola virus VP35 protein is a suppressor of RNA silencing. PLoS Pathog, 2007. 3(6): p. e86
- Chao, J.A., et al., Dual modes of RNA-silencing suppression by Flock House virus protein B2. Nat Struct Mol Biol, 2005. 12(11): p. 952-7
- Gatignol, A., S. Laine, and G. Clerzius, Dual role of TRBP in HIV replication and RNA interference: viral diversion of a cellular pathway or evasion from antiviral immunity? Retrovirology, 2005. 2: p. 65
- Bennasser, Y., M.L. Yeung, and K.T. Jeang, HIV-1 TAR RNA subverts RNA interference in transfected cells through sequestration of TAR RNA-binding protein, TRBP. J Biol Chem, 2006. 281(38): p. 27674-8
- Bennasser, Y., et al., Evidence that HIV-1 encodes an siRNA and a suppressor of RNA silencing. Immunity, 2005. 22(5): p. 607-19
- Bennasser, Y. and K.T. Jeang, HIV-1 Tat interaction with Dicer: requirement for RNA. Retrovirology, 2006. 3: p. 95
- Qian, S., et al., HIV-1 Tat RNA silencing suppressor activity is conserved across kingdoms and counteracts translational repression of HIV-1. Proc Natl Acad Sci U S A, 2009. 106(2): p. 605-10
- Triboulet, R., et al., Suppression of microRNA-silencing pathway by HIV-1 during virus replication. Science, 2007. 315(5818): p. 1579-82
- Christensen, H.S., et al., Small interfering RNAs against the TAR RNA binding protein, TRBP, a Dicer cofactor, inhibit human immunodeficiency virus type 1 long terminal repeat expression and viral production. J Virol, 2007. 81(10): p. 5121-31
- Lin, J. and B.R. Cullen, Analysis of the interaction of primate retroviruses with the human RNA interference machinery. J Virol, 2007. 81(22): p. 12218-26
- Sanghvi, V.R. and L.F. Steel, A Re-Examination of Global Suppression of RNA Interference by HIV-1. PLoS One, 2011. 6(2): p. e17246
- Nathans, R., et al., Cellular microRNA and P bodies modulate host-HIV-1 interactions. Mol Cell, 2009. 34(6): p. 696-709
- Yeung, M.L., et al., Changes in microRNA expression profiles in HIV-1-transfected human cells. Retrovirology, 2005. 2: p. 81
- Chable-Bessia, C., et al., Suppression of HIV-1 replication by microRNA effectors. Retrovirology, 2009. 6: p. 26
- Houzet, L., et al., MicroRNA profile changes in human immunodeficiency virus type 1 (HIV-1) seropositive individuals. Retrovirology, 2008. 5: p. 118
About the Author
Professor John Rossi received his PhD in Genetics from the University of Connecticut in 1976 under the guidance of Dr. Claire Berg and went on to do postdoctoral research in molecular genetics at Brown University under Dr. Arthur Landay. In 1980, he joined the faculty at the City of Hope, and has established a career in studying the biological function and applications of small RNAs. His work with ribozymes led to the first clinical trial of ribozymes for the treatment of HIV-1 infection, and recently has carried our research with three small RNA inhibitors of HIV-1 including a small interfering RNA that has led to a second clinical trial for HIV-1. His work on small RNA mechanisms and applications has resulted in over 250 peer reviewed publications and numerous book chapters and invited commentaries. His lab continues to focus on small RNA biology and applications with an emphasis on applications for HIV and cancer.
Email: [email protected]




