Application of flow cytometry in drug discovery
Posted: 16 December 2010 |
Flow cytometry can be used to advance our understanding of diseases in multiple ways. Drug effects and dosages can be ascertained in vitro, along with patient selection based on mutations and antigen profiles. Within the Diagnostic Biomarkers group of Translational Research at Pfizer, we are utilising flow cytometry in conjunction with other diagnostic tools to assist in gaining a clearer understanding of drug target biology and to identify patients that would benefit most from a specified drug regimen.
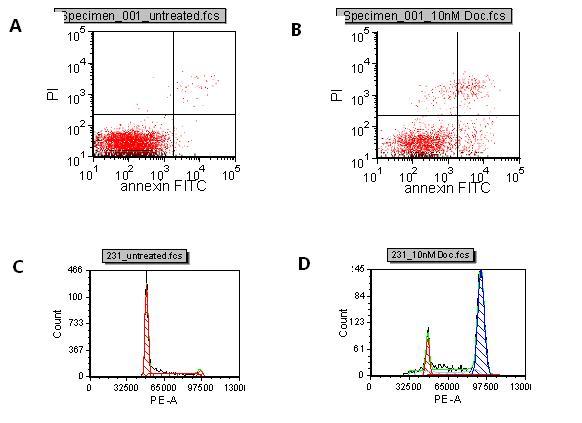

Figure 1 A: Control B: Docetaxel treated Induction of apoptosis with Docetaxel compared to untreated. The lower right quadrant represents cells in early apoptosis and cells in the upper right quadrant are cells in late apoptosis due to PI entering the cells through leaky membranes. C: Control D: Docetaxel treated Cell cycle of control cells and Docetaxel treated cells. The first peak shows cells in G0/G1 and the second peak is cells in G2/M. The breast cancer cell line, MDA MB231, was treated with 10nM docetaxel or DMSO control for 72 hours. Cells were harvested, the nuclei isolated and stained with PI. Samples were run on the LSR II flow cytometer and analyed in FCS Express software. Docetaxel arrests the cells in G2/M
Flow cytometry can be used to advance our understanding of diseases in multiple ways. Drug effects and dosages can be ascertained in vitro, along with patient selection based on mutations and antigen profiles. Within the Diagnostic Biomarkers group of Translational Research at Pfizer, we are utilising flow cytometry in conjunction with other diagnostic tools to assist in gaining a clearer understanding of drug target biology and to identify patients that would benefit most from a specified drug regimen.
In this article, I will discuss how drug effects are followed using cell cycle analysis and apoptotic markers, determine the mechanism of action using a phagocytosis assay and detect tumour metastasis using tumour-specific markers. Additionally, I will discuss how the identification of compounds that act on ‘cancer stem cells’ (CSCs) versus the tumour mass can reduce patient relapse.
Monitoring cell lines in vitro for induction of apoptosis and cell cycle arrest after compound exposure can help identify the in vivo models that will have a higher probability of success.
To assist in the selection of in vivo models, flow cytometry can be used to monitor compound effects on in vitro cell lines for induction of apoptosis and cell cycle arrest. In the early stages of apoptosis, intracellular phosphatidyl serine moves to the surface of the cell allowing them to be detected with a fluorescently labelled annexin V which binds to the phosphatidyl serine with high affinity. In the late stages of apoptosis, the membranes of dying cells become permeable. Propidium iodide (PI), a DNA binding dye, can enter dying cells through their leaky membranes and bind their DNA. By co-staining cells with annexin V and PI, it is possible to differentiate between cells that are in early and late phases of apoptosis (Figure 1, A,B,). Cells in G0 or G1 phases of the cell cycle have two copies of DNA, whereas cells in G2 or M phases have four copies of DNA. PI staining enables the stage of the cell cycle to be determined by measuring the amount of PI bound to the DNA in permeabilised cells or isolated nuclei (Figure 1, C,D,).
Using these simple assays, cell lines that have a greater response to a drug can be easily identified. In addition, the minimum dose needed to arrest the cells in one of the stages of the cell cycle as well as the maximal tolerated dose can be determined. Flow cytometry within the drug discovery paradigm enables us to quickly observe and efficiently triage valuable in vivo models.


Figure 1 A: Control B: Docetaxel treated Induction of apoptosis with Docetaxel compared to untreated. The lower right quadrant represents cells in early apoptosis and cells in the upper right quadrant are cells in late apoptosis due to PI entering the cells through leaky membranes. C: Control D: Docetaxel treated Cell cycle of control cells and Docetaxel treated cells. The first peak shows cells in G0/G1 and the second peak is cells in G2/M. The breast cancer cell line, MDA MB231, was treated with 10nM docetaxel or DMSO control for 72 hours. Cells were harvested, the nuclei isolated and stained with PI. Samples were run on the LSR II flow cytometer and analyed in FCS Express software. Docetaxel arrests the cells in G2/M
Detecting phagocytosis of antibody treated cells
When evaluating biotherapeutics such as humanised antibodies, it is essential to know the mechanism of action. Many antibodies exert their therapeutic effects by binding to overexpressed proteins on a target cell and trigger antibody dependent cellular cytotoxicity (ADCC) or antibody dependent cellular phagocytosis (ADCP). These are a few of the components of the immune system that eliminate immune complexes.
Macrophages, for example, have been shown to be involved in ADCP of the Ramos cell line (Human Burkitt’s Lymphoma) bound to Rituxan. At Pfizer, they have also been shown to be involved in ADCP of the U87 cell line (human glioblastoma) bound to Pfizer’s biotherapeutic, PF5412, which is a dual function fully human IgG1 monoclonal antibody against alpha 5 beta 1.
Monocytes were isolated from normal donor blood using Miltenyi CD14 labelled magnetic beads. They were differentiated into macrophages by treatment with GM-CSF for five days grown in Teflon flasks. The differentiated macrophages and target cells were labelled with different dyes to ensure they were distinguishable population subsets. The macrophages (effector cells) were labelled with PKH26, a membrane dye, and the target cells were labelled with CMFDA, a cytosolic dye. After pre-incubating the biotherapeutic antibody with the target cells, the effector and target cells are mixed together for various time points, and then evaluated on the flow cytometer. Both dyes found together indicate that the macrophage has either bound the target cell or engulfed it.
After four hours, the macrophages have taken up a small proportion of the Rituxan bound target cells (Figure 2). Over time, the double positive population will increase followed by a decrease in the number of Ramos cells as they are digested by the macrophages. U87s bound to PF5412, are taken up specifically by the macrophages as compared to the control KLH antibody (Figure 3. The effect increases in a dose dependent manner (data not shown). This assay provides quantitative data without the use of radioisotopes.
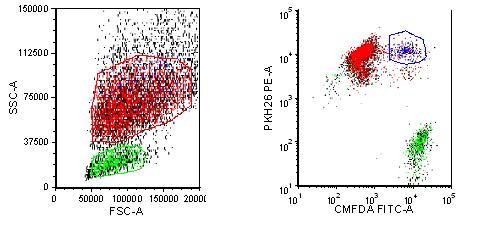

Figure 2 Phagocytosis of Rituxan labeled Ramos cells Macrophages are labelled with PKH26 and Ramos cells are labelled with CMFDA. Rituxan is bound to labelled Ramos cells and then incubated with the labelled macrophages for four hours. The cells are run on an LSR II flow cytometer and analysed with FCS Express software. Dual staining represents either Ramos cells bound to the macrophages through the Fc receptor or the Ramos cells engulfed by the macrophages
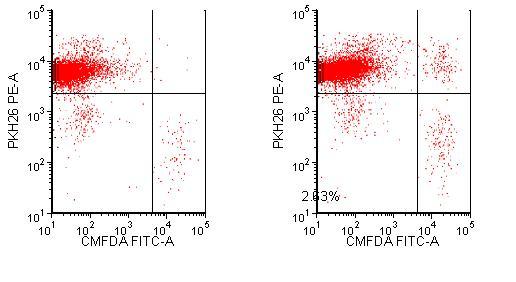

Figure 3 Phagocytosis of biotherapeutic PF5412. Macrophages are labelled with PKH26 and U87s are labelled with CMFDA. U87s are stained with KLH or PF5412 then incubated with the labelled macrophages for four hours. The samples are run on an LSR II flow cytometer and analysed with FCS Express software. The lower right quadrant represents U87 cells that are not bound or engulfed by the macrophages. The upper right quadrant shows U87 bound or phagocytosed by the macrophages
Using flow cytometry to monitor tumour metastasis
Tumour cells often have a unique protein expression pattern, like a fingerprint. It is anticipated that small numbers of metastatic tumour cells can be detected in a patient’s blood due to their expression profile. This can be validated by spiking cancer cells into blood and then staining with antibodies specific to the tumour cells. In this example, 1,000 Raji cells (B cell lymphoma) were spiked into 0.5 millilitres of blood from a nude mouse, and then stained with a CD19 antibody. After collection of 200,000 cells on the flow cytometer, 550 Raji cells could be detected (Figure 4).
The number of metastatic tumour cells in a patient’s blood is usually very small. To detect this small fraction of tumour cells, enrichment techniques can be used to eliminate blood cells that do not co-express the same markers as the tumour cells. This can be accomplished using magnetic beads that are designed to bind to lineage markers on differentiated blood cells. Blood cells that express the lineage markers will be removed, thereby enriching the circulating tumour cell populations. This technique can also be used with in vivo models to determine if induced tumours have become metastatic. Ideally, this approach will be used routinely with cancer patients. By detecting the circulating tumour cells at an early stage, a metastasis can potentially be treated before it takes up residence in the lymph nodes or bone marrow.
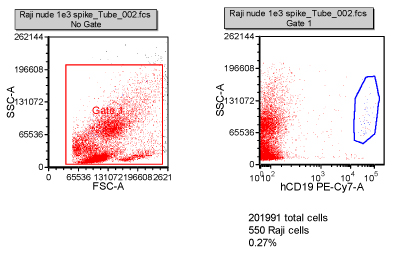

Figure 4: Validation of the detection of circulating tumour cells. One thousand Raji cells were spiked into 0.5ml of nude mouse blood. Using a CD19 antibody specific for the Raji cells, they were detected in the mouse blood at ~0.2 per cent. Two hundred thousand cells were collected and 550 of the Raji cells were distinguished by specific staining
Targeting cancer stem cells
Cancer stem cells (CSCs) or tumour initiating cells (TICs) are a small subset of cells within a tumour that often evade cancer therapies using multiple resistance pathways. There is some evidence that these cancer stem cells may be the cause of the re-emergence of some tumours after treatment. Most treatments kill the primary tumour cells, but the cancer stem cells escape death. Over time, the cancer stem cells differentiate and the tumour develops once more.
The origin of the cancer stem cell appears to come from the loss of a stem cell’s ability to self-regulate. This can be induced by a mutation in various signalling pathways involved in self renewal. CSCs can be identified based on the expression of a variety of markers which are dependent on tumour origin. We screen multiple markers associated with different types of cancer stem cells, following their expression with and without treatment.
Using seven colour panels allows us to see subtle changes in many different markers and gives us the ability to determine which antigens are of interest in a particular model. For example, breast cancer stem cells have been shown to be CD44+/CD24-/lin-, however, there are many other markers, such as EpCam, CD133 and CD166 that might be affected by drug treatment. We have also found that drug resistant tumour cell lines have a different CSC marker profile than the parental line (Figure 5). The marker differences resulting from drug effects and/or drug resistance will be ascertained with further in-depth experiments. These cells will be sorted and tested in other assays to help elucidate the meaning and significance of these changes.
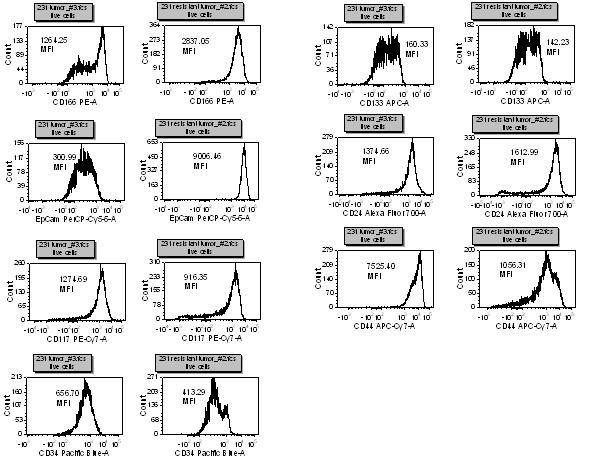

Figure 5. This is a seven antibody panel that detects differences in expression of some ‘cancer stem cell’ markers with Taxol resistant and non-resistant breast tumour cell lines. All of the markers, except CD133, have a different expression profile between the cell types. After staining for dead cells with a fixable dye, non-specific binding is blocked and then optimally titrated antibodies are added to the cells for thirty minutes on ice. The cells are washed thoroughly and then fixed. Samples are then run on an LSR II flow cytometer and analysed with FCS Express software.
Using flow cytometry with in vitro and in vivo studies can help us to further our understanding of the processes involved in drug targets. Using cell cycle and apoptosis studies in vitro allow us to determine which in vivo models will have a better chance of success and provide us with an initial dosage. Phagocytosis studies can be done by a simple flow cytometry assay which can reduce assay time. The effector and target cells can be labelled in multiple colours allowing discrimination of each population. Flow cytometry can be used as an early indicator of metastasis. Proteins that are differentially expressed on the tumour cells can be used to identify the metastatic cells in a patient’s blood, increasing the chance to treat the patient earlier. Cancer stem cells are a relatively new area of cancer research. As our understanding of these rare cells increases, we will be able to find new regimens to target them specifically. In combination with the standard of care, we anticipate a decrease in patient relapse.
Flow cytometry in combination with other biological assays can enhance our understanding of a drug’s impact on tumour cells. While in vitro assays provide a reference for optimal in vivo compound concentrations, flow cytometry can serve to guide strategic experimental and project planning assessments. Knowledge of cellular responses in early-stage drug discovery greatly enables tasks ranging from categorising compound effects to efficiently navigating strategic movement toward in vivo studies. Flow cytometry is a valuable tool that allows us to better understand cancer biology and the effects drugs can have on patients.
References
O’Neill et al AJP Sept 2007, Vol. 171, No. 3. Notch2 signaling induces apoptosis and inhibits human MDA MB231 xenograft growth
Lazar et al PNAS Mar. 14, 2006, Vol. 103, No. 11. Engineered antibody Fc variants with enhanced effector function
Hu et al Cytometry Part A 77A:213-219, 2010. Detection of circulating tumor cells in breast cancer patients utilizing multiparameter flow cytometry and assessment of the prognosis of patients in different CTCs levels
Kakarala et al JCO Jun 2008, vol. 26, No. 17. Implications of the cancer stem cell hypothesis for breast cancer prevention and therapy
About the Author
Dana Buckman
Dana has been a flow cytometry specialist since 1992 working in industry, focusing on cancer immunology. Dana is currently a senior scientist at Pfizer-La Jolla in the biomarkers group of translational research.




