RNAi screening in the era of high-throughput genetics
Posted: 12 December 2009 | Steven Haney, Associate Fellow, Pfizer | No comments yet
The use of RNAi screening to identify potential drug targets has enjoyed great success in recent years as a robust method for linking genes to a disease process through a functional assessment of a gene in an experimental model1. True, RNAi screening is complicated by problems such as off-target effects and toxicities associated with various properties of RNAi molecules, but effort from many groups has produced a coherent set of guidelines that are useful for most RNAi screens2-4. As such, large scale RNAi screens can be performed by moderately-resourced laboratories. RNAi screens have identified genes involved in resistance to anticancer chemotherapeutics, viral and bacterial infection and specific signal transduction pathways. Now that the technical complications have been identified and largely mitigated, RNAi screening has joined other genomic technologies, such as transcriptional profiling, as a method for broadly surveying the human genome for roles in a given disease or a response to a treatment.
The use of RNAi screening to identify potential drug targets has enjoyed great success in recent years as a robust method for linking genes to a disease process through a functional assessment of a gene in an experimental model1. True, RNAi screening is complicated by problems such as off-target effects and toxicities associated with various properties of RNAi molecules, but effort from many groups has produced a coherent set of guidelines that are useful for most RNAi screens2-4. As such, large scale RNAi screens can be performed by moderately-resourced laboratories. RNAi screens have identified genes involved in resistance to anticancer chemotherapeutics, viral and bacterial infection and specific signal transduction pathways. Now that the technical complications have been identified and largely mitigated, RNAi screening has joined other genomic technologies, such as transcriptional profiling, as a method for broadly surveying the human genome for roles in a given disease or a response to a treatment.
The use of RNAi screening to identify potential drug targets has enjoyed great success in recent years as a robust method for linking genes to a disease process through a functional assessment of a gene in an experimental model1. True, RNAi screening is complicated by problems such as off-target effects and toxicities associated with various properties of RNAi molecules, but effort from many groups has produced a coherent set of guidelines that are useful for most RNAi screens2-4. As such, large scale RNAi screens can be performed by moderately-resourced laboratories. RNAi screens have identified genes involved in resistance to anticancer chemotherapeutics, viral and bacterial infection and specific signal transduction pathways. Now that the technical complications have been identified and largely mitigated, RNAi screening has joined other genomic technologies, such as transcriptional profiling, as a method for broadly surveying the human genome for roles in a given disease or a response to a treatment.
Genomic technologies report on differences between states, such as normal and transformed cells or tissues, and how cells respond to a perturbation. Each technology has inherent strengths and weaknesses. Effectively using these approaches is rooted in understanding how to combine them such that the strengths of one mitigate the weaknesses in another. Transcriptional profiling reports on genes that are differentially regulated, but can not distinguish between those that cause a phenotypic change from those that result from or define the phenotypic change. Protein-protein interaction networks identify functional groups of proteins (and their cognate genes) that can be combined with transcriptional profiling data to establish network-level changes. Thus, an integrated approach to unbiased data sets can reinforce the findings of individual studies, and can enable specific molecular hypotheses to be developed and tested5. Since transcriptional profiling experiments typically hundreds, or even thousands, of genes this has been a daunting challenge. In contrast, RNAi screening is valuable for identifying genes relevant to a biological process through a functional consequence and complements methods that are agnostic to cause-or-effect differences.
More recently, human genetics has accelerated its pace as a method for identifying differences between healthy and diseased states, and by extension, differences between normal signaling pathways and those that are dysfunctional. These pathways are potentially sensitive to therapeutic intervention. Most methods have improved as a result of the human genome sequencing efforts, either in the data generated, such as high density SNP (single nucleotide polymorphism) chips, or in the methods that had been developed during the program, such as Next Generation Sequencing (NGS). SNP chips have been used to track the prevalence of common polymorphisms (generally defined as occurring in >5% of the population) in normal and disease groups. In genome-wide association (GWA) studies, SNPs showing an enrichment of one allele in one of the study groups can implicate one or more genes within the region of that SNP that may play a role in the disease. NGS can identify mutations in coding regions of genes directly.
The advantage of leveraging human genetic data is that the genes identified are defined by their role in human disease. In contrast, RNAi screening or transcriptionally profiling of a human cancer cell line identifies genes that are relevant to the survival of that cancer line under the conditions tested (typically as a mono-culture in atmospheric oxygen and enriched media), but translating such effects to cancer in humans requires that the model system truly recapitulate events in the tumour. Such caveats can be overstated, for example breast cancer models of Her-2NEU over-expression are effective for evaluating anti-Her-2NEU therapeutics, despite being cultured ex vivo for decades. However, an initiative will only succeed in developing a therapeutic if a concordance between a model and a disease mechanism exists. The strongest advantage to human genetic studies of disease is that this concordance is explicit. Genes or chromosomal loci are only identified as significant if they are present to different degrees between the control and the disease groups. Therefore, a gene that is identified directly (by NGS) or implicated by its locus (through SNP genotyping) affects the onset or progression of the disease under study in an authentic human population. While not directly related to the methodology, an additional advantage to SNP-based studies is that most studies identify only a limited number of genes, often less than 20 genes and frequently less than 10. In contrast, transcriptional profiling and RNAi screening typically identify many more genes, and these gene lists are known to include many that are responding to, rather than driving, the biological problem (in the case of transcriptional profiling) or to include technical artifacts (false positives in the case of RNAi screening). Larger scale sequencing methods are also generating larger lists of candidate genes, but informatics methods are developing in parallel to reduce the total number of mutations to a refined list of candidates (discussed below).
Understanding why the mutations confer sensitivity to a disease is the major challenge. If genes identified in genetic studies are (indeed) defined by the statistical significance, there is frequently little information available on why the mutation, or the gene affected by the mutation, plays such a causative role. In fact, no information can be inferred on what the relevant cell type, tissue or organ is. Additional methods are necessary to develop a mechanistic context for understanding how the mutation affects disease. Expression studies are valuable for defining a list of cell types or tissues to focus on, but even then caution is needed if the expression studies are themselves generated in cell lines or primary culture of cells taken out of their native environment. Ironically, well-characterised genes can lead to prematurely focusing on a particular cell type, when in fact, several cell types ought to be considered.
Genes identified through high-throughput genetics
An overview of recent studies in human genetics has been presented above, but it is worth taking a moment to discuss the results generated from them in more detail. Sequencing methods have focused on pre-selected genes, and so they have not been completely unbiased, but the number of genes has been increasing, and sequencing the exome (all exons) is now possible6. Mutations identified by sequencing are still evaluated for their statistical significance, based on their occurrence in a disease group and compared to the frequency of occurrence in a control group, so even though coding sequence mutations are strongly indicative of a consequential impact on disease, such mutations are not truly ‘smoking guns,’ and still require additional validation. In the case of somatic mutations found in human tumours, many mutations are identified in the tumour samples, and the oncogenic (driver) mutations need to be distinguished from other (passenger) mutations. Exon sequencing is distinct from transcriptome sequencing, in that the DNA is sequenced rather than RNA, and therefore expression information can not be derived from the mutational data.
SNP studies have grown from early work cataloging normal genetic variation in human populations, and then searching for unusual patterns (distributions) of specific variants that correlate with the occurrence of a disease or another measurable parameter. Each loci identified in such a study may implicate one or more genes. SNPs identified as most significant are compared with additional SNPs and the genes that reside in the locus. In the example shown in Figure 1, the most significant SNP identified in the figure, rs10923931, is shown as a diamond near the top of the figure. Additional SNPs are also shown, those that correlate with query SNP are shown in darker colour than those that do not. The recombination rate is shown as the blue line, chromosomal regions that lie within the recombination boundaries are associated with the query SNP, and in this example include the genes NOTCH2 and ADAM30. Those that lie across the boundary are not associated with the query SNP, such as the genes HMGC182, PGDH3 and REG4.
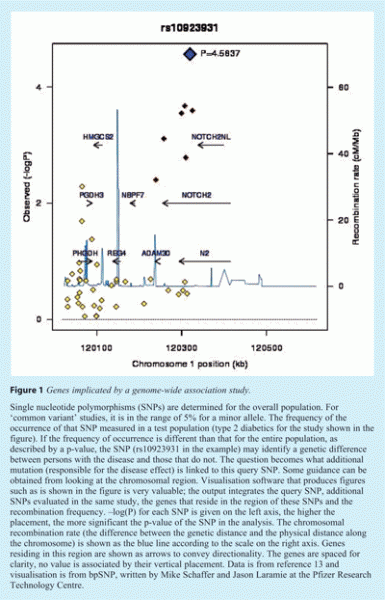

Genes identified in these studies include a few that are very likely to play some role in the disease in question, although it is most likely that only one gene within a locus is actually playing such a role. The challenge then becomes determining the function of the gene as it pertains to the disease. This is particularly difficult when one considers the genes that are identified in these studies. Looking as several GWA studies, the genes considered as candidates include some that can easily be fit into a mechanistic rationale while others are poorly characterised or implicate biological processes not normally associated with the disease. A couple of examples are listed in Table 1. In many cases, fundamentally novel genes are identified in addition to genes that have been well-characterised in at least one cell type. As such, some genes may present a path forward for their study, whereas others have no known function or pathway. Additionally, our understanding of the genome is still evolving, and alternative transcripts exist that are not targeted in most RNAi studies. One example is ARNIL, an expressed transcript within the CDKN2B locus. Variants that map to this locus are associated with diabetes7 and atherosclerosis8. This region includes a long transcript (designated ARNIL) in the opposite orientation to CDKN2B and spans many of the SNPs showing a correlation with these diseases. Disease-associated variants have been shown to affect expression CDKN2A, CDKN2B and ARNIL, all of which lie within this region9. Therefore, a strict interpretation of the available data is that any of the three genes within the region identified in several studies may be playing a causative role, and it is not necessarily true that the same mutation or gene is causing multiple diseases.
Lastly, while the disease-causing mutations have occasionally been identified as coding changes to the open reading frame (for example some of the MODY group of diabetes genes that include TCF7L2 and GCK), in many cases such mutations have not been identified. The mutations that cause disease are likely to alter expression levels. This is an important observation for validation studies, as it is consistent with the use of RNAi as a validation method, since it explicitly affects gene expression levels. If a common mechanism of increased disease risk is not through the altered function of the gene product, but instead is through altered expression of the gene product, the use RNAi screening to investigate the role of genes in disease is more than just a surrogate method of perturbation, it can reasonably model the authentic effect of the mutation.
Validation of novel and uncharacterised genes for roles in disease
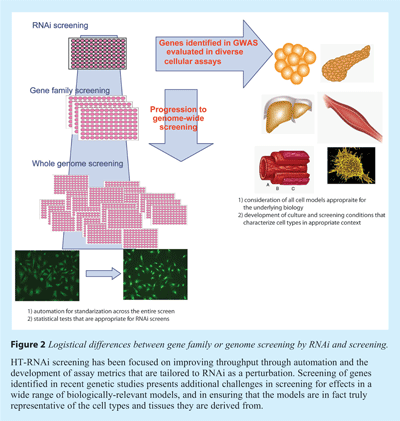
Since a genetic mutation identifies physical region of a chromosome, and does not directly implicate a gene or even the cell type where the mutation exerts its effect, a full consideration of the candidate targets requires broadening list of genes screened, as well as the biological systems available to study these genes. Some of the factors that need to be adjusted to broaden the biological systems are diagramed in Figure 2. RNAi screening in support of GWA results is distinct from genome-scale RNAi screening in several important ways. A couple of these are a result of the differences in scale of screening for the two approaches. Screening a family of druggable targets or a larger group of genes requires standardisation of the screening assays and data analysis10. For example, the list of protein kinases approaches a thousand genes and this group is frequently screened with additional groups of druggable targets, such as the G-protein coupled receptors (GPCRs) or the nuclear hormone receptors (such as the glucacorticoid receptor). Configurations of the druggable targets gene families frequently total more than 5000 genes. Successfully screening such a library requires robust statistical methods that are appropriate for RNAi screens2,11, and follow-on methods to eliminate false positives, which are frequent in large scale screens3,4. Screening at a large scale means that several concessions are important. For example, the cells or cell lines used must be amenable to handling with automation, including sitting in reservoirs for periods of time, and must plate reproducibly. As a consequence, large scale RNAi screening requires changes in the cellular systems to accommodate screening. Most commonly, this includes immortalisation or transformation. These changes eliminate some biological questions from being addressed by large scale screening, however screens to characterise specific signaling pathways (e.g. JAK-STAT, EGFR) and some cellular processes (e.g. pathways used by infectious agents to invade a host cell) have been successful.


In contrast, screening for a causative role by an uncharacterised gene increases the necessity that the cell system used be as close to its native environment as possible. In large part, this is because a causative role may still be indirect, such as cell stress affecting glucose regulation in the liver or skeletal muscle, and such roles can not be determined a priori. This applies both to the genetic alterations in the cell system screened (an immortalised line will have lost expression of CDKN2A and CDKN2B, so these genes, as well as genes that impinge on these, will fail to show an effect in a functional test), as well as to the culture of the cells themselves. Primary cells are challenging to work with, and minor changes in their handling can cause critical cell type-specific functions to be lost. These effects have been noted by work in the toxicology and stem cell biology fields for many cell types. Therefore, many potential biological functions can not be determined through screening at a large scale, and a focused screen with a carefully managed cell model is essential. This is important because it is likely that any given mutation or gene knockdown will eventually produce a phenotype or change in a signal transduction pathway. When such a change resulting from a mutation is identified, it may be tempting to conclude that the role for the mutation has been established, but it is only a role for the mutation in a cellular system, and may not be a role for the mutation in disease.

Concluding remarks
RNAi screening in a wide range of cellular systems to establish a disease mechanism is both subject to the technical concerns intrinsic to RNAi screening in general, such as off-target effects, as well as a strong link to biological relevance, because the observance of an effect in the model system does more than implicate a gene in a pathway, it is intended as evidence of an effect on disease. An example of this difference would be seen if a siRNA causes a change in assays in more than one system. If the disease area was diabetes, an effect could be observed in models for pancreatic beta-cells, skeletal muscle and the liver. An effect in all three systems would provide three models for studying the role of the gene in a biological context, but it may be only one of these systems that actually produces the effect in clinical diabetes. If the study looked at a genetic basis for an increased likelihood of cancer, effects in stromal, endothelial and inflammatory cells could similarly signify more contexts to study the gene than are relevant to the disease. While the characterisation of disease has been genuinely advanced through the application of recently developed methods for genetic analysis, determining the role of these mutations has also identified gaps in the linkage between many biological models used to develop therapeutics and the authentic mechanisms of human disease.
Acknowledgement
The SNP described in Figure 1 on page 40 makes use of data generated by the Wellcome Trust Case-Control Consortium. A full list of the investigators who contributed to the generation of the data is available from www.wtccc.org.uk. Funding for the project was provided by the Wellcome Trust under award 076113.
References
- Boutros, M., and J. Ahringer. 2008. The art and design of genetic screens: RNA interference. Nature Reviews Genetics 9:554-566.
- Birmingham, A., L. M. Selfors, T. Forster, D. Wrobel, C. J. Kennedy, E. Shanks, J. Santoyo-Lopez, D. J. Dunican, A. Long, D. Kelleher, Q. Smith, R. L. Beijersbergen, P. Ghazal, and C. E. Shamu. 2009. Statistical methods for analysis of high-throughput RNA interference screens. Nature Methods 6:569-575.
- Echeverri, C. J., P. A. Beachy, B. Baum, M. Boutros, F. Buchholz, S. K. Chanda, J. Downward, J. Ellenberg, A. G. Fraser, N. Hacohen, W. C. Hahn, A. L. Jackson, A. Kiger, P. S. Linsley, L. Lum, Y. Ma, B. Mathey-Prevot, D. E. Root, D. M. Sabatini, J. Taipale, N. Perrimon, and R. Bernards. 2006. Minimizing the risk of reporting false positives in large-scale RNAi screens. Nature Methods 3:777-779.
- Haney, S. A. 2007. Increasing the robustness and validity of RNAi screens. Pharmacogenomics 8:1037-1049.
- Lum, P. Y., J. M. Derry, and E. E. Schadt. 2009. Integrative genomics and drug development. Pharmacogenomics 10:203-212.
- Ng, S. B., E. H. Turner, P. D. Robertson, S. D. Flygare, A. W. Bigham, C. Lee, T. Shaffer, M. Wong, A. Bhattacharjee, E. E. Eichler, M. Bamshad, D. A. Nickerson, and J. Shendure. 2009. Targeted capture and massively parallel sequencing of 12 human exomes. Nature 461:272-276.
- Lyssenko, V., A. Jonsson, P. Almgren, N. Pulizzi, B. Isomaa, T. Tuomi, G. Berglund, D. Altschuler, P. Nilson, and L. Groop. 2008. Clinical risk factors, DNA varients and the development of type 2 diabetes. New England Journal of Medicine 359:2220-2232.
- Helgadottir, A., G. Thorleifsson, A. Manolescu, S. Gretarsdottir, T. Blondal, A. Jonasdottir, A. Jonasdottir, A. Sigurdsson, A. Baker, A. Palsson, G. Masson, D. F. Gudbjartsson, K. P. Magnusson, K. Andersen, L. A.I., V. M. Backman, S. Matthiasdottir, T. Jonsdottir, S. Palsson, H. Einarsdottir, S. Gunnarsdottir, A. Gylfason, V. Vaccarino, W. C. Hooper, M. P. Reilly, C. B. Granger, H. Austin, D. J. Rader, S. H. Shah, A. A. Quyyumi, J. R. Gulcher, G. Thorgeirsson, U. Thorsteinsdottir, A. Kong, and K. Stefansson. 2007. A common variant on chromosome 9p21 affects the risk of myocardial infarction. Science 316:1491-1493.
- Liu, Y., H. K. Sanoff, H. Cho, C. E. Burd, C. Torrice, K. L. Mohlke, J. G. Ibrahim, N. E. Thomas, and N. E. Sharpless. 2009. INK4/ARF transcript expression is associated with chromosome 9p21 variants linked to atherosclerosis. PLoS One 4:e5027.
- Echeverri, C. J., and N. Perrimon. 2006. High-throughput RNAi screening in cultured cells: a user’s guide. Nature Reviews Genetics 7:373-384.
- Zhang, X. D. 2007. A pair on new statistical parameters for quality control in RNA interference high-throughput screening assays. Genomics 89:552-561.
- Houlston, R. S., E. Webb, P. Broderick, A. M. Pittman, M. C. Di Bernardo, S. Lubbe, I. Chandler, J. Vijayakrishnan, K. Sullivan, S. Penegar, C. C. A. S. Consortium, L. Carvajal-Carmona, K. Howarth, E. Jaeger, S. L. Spain, A. Walther, E. Barclay, L. Martin, M. Gorman, E. Domingo, A. S. Teixeira, C. Consortium., D. Kerr, J. B. Cazier, I. Niittymäki, S. Tuupanen, A. Karhu, L. A. Aaltonen, I. P. Tomlinson, S. M. Farrington, A. Tenesa, J. G. Prendergast, R. A. Barnetson, R. Cetnarskyj, M. E. Porteous, P. D. Pharoah, T. Koessler, J. Hampe, S. Buch, C. Schafmayer, J. Tepel, S. Schreiber, H. Völzke, J. Chang-Claude, M. Hoffmeister, H. Brenner, B. W. Zanke, A. Montpetit, T. J. Hudson, S. Gallinger, I. C. C. G. A. Consortium, H. Campbell, and M. G. Dunlop. 2008. Meta-analysis of genome-wide association data identifies four new susceptibility loci for colorectal cancer. Nature Genetics 40:1426-1435.
- Consortium, T. W. T. C. C. 2007. Genome-wide association study of 14,000 cases of seven common diseases and 3000 shared controls. Nature 447:661-678.




