DNA microarrays and multidrug resistant bacteria
Posted: 1 March 2018 | Dr Payam Behzadi | No comments yet
In parallel with the use of a wide range of antibiotics for treating infectious diseases worldwide, antimicrobial resistance has significantly increased. For this reason, the application of an advanced pangenomic technique like DNA microarray can be an effective, rapid, accurate, sensitive, specific and flexible methodology for detecting, identifying and determining antimicrobial-resistant bacteria.


In the ancient world, including Ancient Persia (Iran – in particular, near the Caspian Sea in the North and the Persian Gulf in the South of Iran’s plateau), plants, medicinal herbs and fungi were used to treat various diseases and infections. Today, in addition to medicinal herbs, plants and fungi, different types of antibiotics taken from bacteria and fungi are used as the main option for medical treatment. Simultaneously, with the beginning of the use of medicinal substances and antibiotics, resistant strains of microorganisms began to emerge.1,2
Antibiotic-resistant bacteria are not a new feature in medicine, but increased antibiotic use leading to multi-drug resistance is now a global healthcare concern. Methicillin-resistant Staphylococcus aureus (MRSA), methicillin-resistant it Staphylococcus epidermidis (MRSE), vancomycin-resistant Enterococci (VRE), and pandrug-resistant bacteria such as Acinetobacter baumannii, Escherichia coli and Pseudomonas aeruginosa, are important common multidrug-resistant bacteria that can cause complicated morbidity and even mortality.3-6
Bacteria possess a wide range of antimicrobial resistance mechanisms against different types of antibiotics, and several microbiological methods can be used to identify multidrug-resistant bacteria. But, this article focuses on DNA microarray technology as an effective methodology for recognising antibiotic-resistance properties among bacterial strains.
A variety of genes can contribute to antibiotic resistance. Genes responsible for biosynthesis pathway alternation, beta-lactamase enzymes and efflux pumps are the most significant. Moreover, mutations in some genes may reduce the entrance of antibiotics into the bacterial cells, which is known to be an effective antimicrobial resistance property. It is clear that both chromosomes and plasmids carry different types of multidrug resistance genes.7,8

Beta-lactams and glycopeptides are drugs which affect bacterial cell wall biosynthesis. Rifampin and fluoroquinolones inhibit RNA and DNA syntheses. Aminoglycosides and macrolides can inhibit protein biosynthesis via different methods. And polymyxins and sulfonamides affect bacterial cells through destruction of cytoplasmic membrane and metabolic pathway inhibition respectively.5,7
Interestingly, some multidrug resistance genes may be acquired through vertical evolution by one or several mutations within chromosomal genes, while other antimicrobial resistance genes may be acquired via plasmids or other mobile genetic elements. The latter, which is known as horizontal evolution, is very common among bacterial strains. There are many multidrug resistance genes which are of great concern for the global healthcare system. Therefore, this article provides an overview of the most common genes of extended-spectrum beta-lactamases (ESBLs), VRE and MRSA.3,5,7
DNA microarray technology
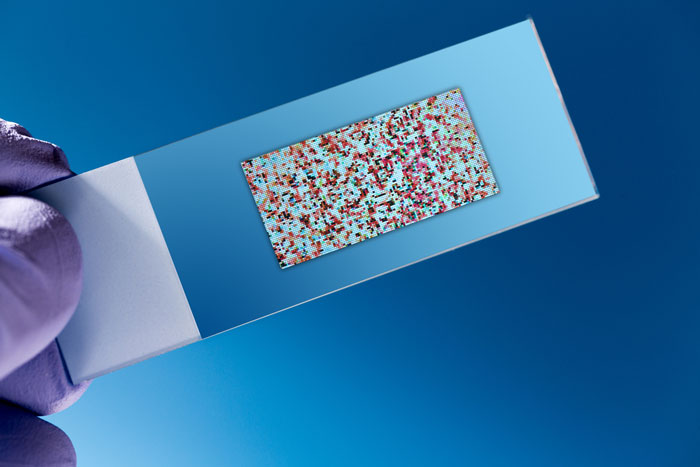
Microarray technology is an advanced pangenomic diagnostic which can be used for detection and identification of different cells and microorganisms. Indeed, this technology gives us an excellent opportunity to identify genes with a high resolution. Microarray as a miniaturised lab-on-a-chip technology is a sensitive, specific, accurate, rapid, reliable and flexible diagnostic tool that enables the detection and identification of infectious and non-infectious diseases, genetic mutations and disorders, and microorganisms, with a high quality. Regarding microorganisms, this technology can be applied to pathogenic and non-pathogenic, antimicrobial sensitive and antimicrobial resistant bacteria, making it flexible.9-12
Applications for DNA microarray technology include gene-profiling preparation and gene expression. Furthermore, to maximise the reliability, flexibility, specificity, sensitivity and accuracy of the outcomes, the use of proper platforms, probe sets, software and tools, is unavoidable. Professional bioinformaticians are also necessary to reduce the risk of bias in final outcomes. Microarray technology provides invaluable qualitative or qualitative outcomes, completely in line with our goal for a scientific investigation.11,13,14
DNA microarray and multidrug-resistant bacteria
There is a range of normal molecular diagnostics, such as polymerase chain reaction (PCR). PCR is a good choice for detecting and identifying a single gene or a limited number of genes simultaneously. But when trying to detect a huge number of genes, such as those involved in antimicrobial resistance, PCR is not a good choice because it will be time-consuming and very expensive. So, this technique must be replaced by microarray technology, which is an economical and proper methodology to detect and identify a vast range of genes simultaneously and quickly.
Various types of multidrug resistance genes must be recognised immediately. Some bacteria, including E.coli, Klebsiella pneumoniae, A. baumannii, P. aeruginosa, Staph. aureus, Staph. epidermidis and enterococci, may have a mass of multidrug-resistance genes.15 Having an up-to-date map in epidemiological studies may help to control improper situations regarding untreated infections and infectious diseases. Among antimicrobial-resistant bacteria, those which include ESBL genes can cause a serious problem during treatment. ESBL genes are easily transferred between Gram-negative enteric bacteria, eg, E. coli and K. pneumoniae via plasmids. Interestingly, there is a diversity of ESBL genes, comprising CTX-M group, SHV and TEM. So, having a rapid and accurate recognition of different types of ESBLs is very important to achieve a definite treatment and preventing the spread of multidrug-resistant strains. The presence of different types of ESBLs is a big concern when it is about enteric bacteria like E. coli and K. pneumoniae important causative agents of urinary tract infections.4,16-22
Vancomycin-resistant bacteria (VRB) involving VRE and VR Staph. aureus (VRSA) are common antimicrobial-resistant bacteria which include transposons bearing vanA genes. In accordance with previous studies, those transposons that bear vancomycin-resistance genes can transfer genes between different types of bacterial genera. Besides, related transposons may bear antimicrobial resistance genes other than vanA. The presence of vanA genes in enterococci is a considerable healthcare problem because the Gram-positive enterococci are resistant to a wide range of antibiotics, involving aminoglycosides, tetracyclines, beta-lactam antibiotics, etc. Therefore, the production of VRE may be a great concern requiring a definite treatment.4,5
Genes which code penicillin-binding proteins (PBPs) may be transferred through transformation. PBP-coding genes are detected in different types of bacteria and, in particular, in Staph. Aureus. MRSA may lead to problems regarding a definite treatment for infections.4,5
Knowing local strains and probable resistance genes in bacterial hosts help us to replace other antibiotics and have a definite treatment. An advanced pangenomic diagnostic technology like microarray is completely dependent on databases, tools and software to provide an accurate, rapid, sensitive, specific, reliable and flexible diagnosis. Microarray technology is divided into two stages: in-silico and in-vitro. The in-silico procedures are known as the ‘dry lab’ section, in which probes are designed. The probe-designing section is the most important part of the technology.
Effective databases, tools and software influence final outcomes, so a professional bioinformatician is essential for flexible and reliable probes. After the probe-designing section, the wet-lab stage is performed to detect and identify probable genes via lab instruments. Using databases like the National Center for Biotechnology Information (NCBI) may help to create complete genome map for designing necessary and effective probes. As mentioned before, precise information about local bacterial strains and the epidemiologic prevalence of antimicrobial resistance genes enables flexible, accurate and rapid diagnostics to substitute an effective antibiotic for a definite treatment.23-25
Biography
Dr Payam Behzadi is Assistant Professor, Department of Microbiology, College of Basic Sciences, Shahr-e-Qods Branch, Islamic Azad University, Tehran, Iran. He has a BSc and MSc in Microbiology and a PhD in Molecular Biology. Dr Behzadi has authored more than 15 academic books and more than 50 original and review articles.
References
- Petrovska BB, Cekovska S. Extracts from the history and medical properties of garlic. Pharmacognosy reviews. 2010;4(7):106.
- Behzadi P, Behzadi E. Modern Industrial Mycology. Tehran: Persian Science & Research Publisher. 2012.
- Behzadi P, Behzadi E, Ranjbar R. Multidrug-Resistant Bacteria. Infectio ro. 2014(39):29.
- Nikaido H. Multidrug resistance in bacteria. Annual review of biochemistry. 2009;78:119-46.
- Tenover FC. Mechanisms of antimicrobial resistance in bacteria. The American journal of medicine. 2006;119(6):S3-S10.
- Behzadi P, Behzadi E, Ranjbar R. Antibiotic therapy for acute otitis media in children. ORL ro. 2014;7(23).
- Behzadi P, Behzadi E. Microbiology of Prokaryotes. 1st ed. Tehran: Eslam Shahr and Shahr-e-Qods Branches, Islamic Azad University. 2006.
- Behzadi P, Behzadi E. Environmental Microbiology. 1st ed. Tehran: Niktab Publisher. 2007.
- Behzadi P, Behzadi E, Ranjbar R. The application of Microarray in Medicine. ORL ro. 2014;7:24.
- Behzadi P, Najafi A, Behzadi E, Ranjbar R. Detection and identification of clinical pathogenic fungi by DNA microarray. Infectioro. 2013;35(3):6-10.
- Behzadi P, Behzadi E, Ranjbar R. Microarray probe set: Biology, bioinformatics and biophysics. Alb Med J. 2015;2:5.
- Behzadi P, Behzadi E, Ranjbar R. Microarray data analysis. Alb Med J. 2014;7:8.
- Behzadi P, Ranjbar R. Microarray long oligo probe designing for Bacteria: An in silico pan-genomic research. Alb Med J. 2016;6(8):9.
- Nsofor CA. DNA microarrays and their applications in medical microbiology. Biotechnology and Molecular Biology Reviews. 2014;9(1):1-11.
- Card R, Zhang J, Das P, Cook C, Woodford N, Anjum MF. Evaluation of an expanded microarray for detecting antibiotic resistance genes in a broad range of gram-negative bacterial pathogens. Antimicrobial agents and chemotherapy. 2013;57(1):458-65.
- Iovleva A, Bonomo RA. The ecology of extended-spectrum β-lactamases (ESBLs) in the developed world. Journal of Travel Medicine. 2017;24(suppl_1):S44-S51.
- Behzadi P, Behzadi E. The microbial agents of urinary tract infections at central laboratory of Dr. Shariati Hospital, Tehran, Iran. Turk Klin Tip Bilim. 2008;28(4):445.
- Jahandeh N, Ranjbar R, Behzadi P, Behzadi E. Uropathogenic Escherichia coli virulence genes: invaluable approaches for designing DNA microarray probes. Central European journal of urology. 2015;68(4):452.
- Ranjbar R, Tabatabaee A, Behzadi P, Kheiri R. Enterobacterial repetitive intergenic consensus polymerase chain reaction (ERIC-PCR) genotyping of escherichia coli strains isolated from different animal stool specimens. Iranian Journal of Pathology. 2017;12(1):25-34.
- Behzadi E, Behzadi P. The role of toll-like receptors (TLRs) in urinary tract infections (UTIs). Central European journal of urology. 2016;69(4):404.
- Behzadi P, Behzadi E, editors. Uropathogenic Escherichia coli: An Ideal Resource for DNA Microarray Probe Designing. International Conference on Bioinformatics and Biomedical Engineering. 2017. Springer.
- Bogner C, Miethke T, Wantia N, Gebhard F, Busch D, Hoffmann R. Differences in ESBL Genes between E. coli, Klebsiella spp. and Enterobacter Cloacae Strains. International Journal of Clinical & Medical Microbiology. 2016.
- Behzadi P, Najafi A, Behzadi E, Ranjbar R. Microarray long oligo probe designing for Escherichia coli: an in-silico DNA marker extraction. Central European journal of urology. 2016;69(1):105.
- Ranjbar R, Behzadi P, Mammina C. Respiratory Tularemia: Francisella Tularensis and Microarray Probe Designing. The Open Microbiology Journal. 2016;10:176.
- Frye JG, Lindsey RL, Rondeau G, Porwollik S, Long F, McClelland M, et al. Development of a DNA microarray to detect antimicrobial resistance genes identified in the National Center for Biotechnology Information database. Microbial Drug Resistance. 2010;16(1):9-19.