Mass spectrometry for gene therapy vector analysis
Posted: 8 October 2021 | Hannah Balfour (European Pharmaceutical Review) | No comments yet
Researchers show combining mass spectrometry and a gas-phase electrophoretic mobility molecular analyser improves detection of full and empty virus-like particles for gene therapies.


Adeno-associated viruses (AAV) are the leading virus-like particle (VLP) vehicles for gene therapies, but how can AAVs either lacking or carrying an engineered genome be identified?
Native mass spectrometry (MS) is known for its ability to gather highly accurate molecular weights without changing the native biological conformation of analytes, but according to experts, once applied to larger biologics (megadalton species), MS becomes challenging and requires customised instruments.
In a new paper published in the Journal of Mass Spectrometry, researchers showed that, by combining native nano-electrospray gas-phase electrophoretic mobility molecular analyser (GEMMA) (native nES GEMMA) and native nano-electrospray ionization quadrupole reflectron time-of-flight mass spectrometry (native nESI QRTOF MS), they could estimate the molecular weight of AAVs to determine which were carrying the engineered genome.
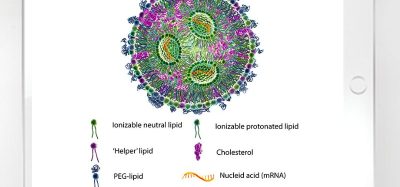
AAVs are a proteinaceous shells derived from adenoviruses, but lacking any viral genomic material, rendering them non-infectious. In gene therapies, these VLPs are used to carry a piece of single-stranded DNA (ssDNA) up to 4.7 kb in size.
In the study researchers created two forms of AAV serotype 8 (AAV8), one “empty”, comprising just the AAV8 shell and the other “filled”, with an engineered genome inside the shell. These were then analysed using native nES GEMMA and nESI QRTOF MS, two technologies adapted for studying protein complexes in their native conformation.
The researchers sought to combine the techniques because both have limitations, nES GEMMA, for example, cannot directly determine the exact molecular weight of particles – instead it provides the dry-surface diameter of particles in the nanometre range and then weight can be inferred using electrophoretic mobility diameter (EMD)/molecular weight correlations for VLPs. Native MS instead generates mass-over-charge results but suffers from low ion transmission efficiency at very high molecular weight and deconvolution challenges. To enable nES GEMMA to be used to help interpret native MS data in the research, the molecular weights were determined from a literature based EMD/molecular weight correlation.
They found that based on EMD/MW correlations, nES GEMMA analysis found the molecular weights of 3670 ± 69 kDa and 4,751 ± 47 kDa for empty and filled AAV8s, respectively. Though able to detect both empty and filled AAV8s, nESI QRTOF MS could not resolve their charges to provide molecular weights. To overcome this, the data inferred by the EMD/MW correlations and the size of the encapsulated genome were used to estimate with high accuracy the number of charges enveloping the AAV8s and thus the molecular weights for each type of VLP.
Using this method, they found that, based on MS data, empty capsids has a molecular weight of 3710 kDa (1.1 percent difference from the GEMMA correlation-based value) and filled VLPs, 5005 kDa (5.4 percent difference from the GEMMA correlation-based value). The researchers said these results corroborate the expected values, underscoring the fidelity of the EMD/molecular weight correlations and that the combination of native nES GEMMA and native nESI QRTOF MS is “very powerful, enabling the in-depth interpretation of data derived from each of these two analysis methods alone to a much higher level of detail”.
Related topics
Analytical techniques, Biologics, Gene therapy, Genomics, Mass Spectrometry, Technology, Therapeutics